Beyond the Data Desert: Advanced Strategies to Overcome Data Scarcity in Molecular Property Prediction
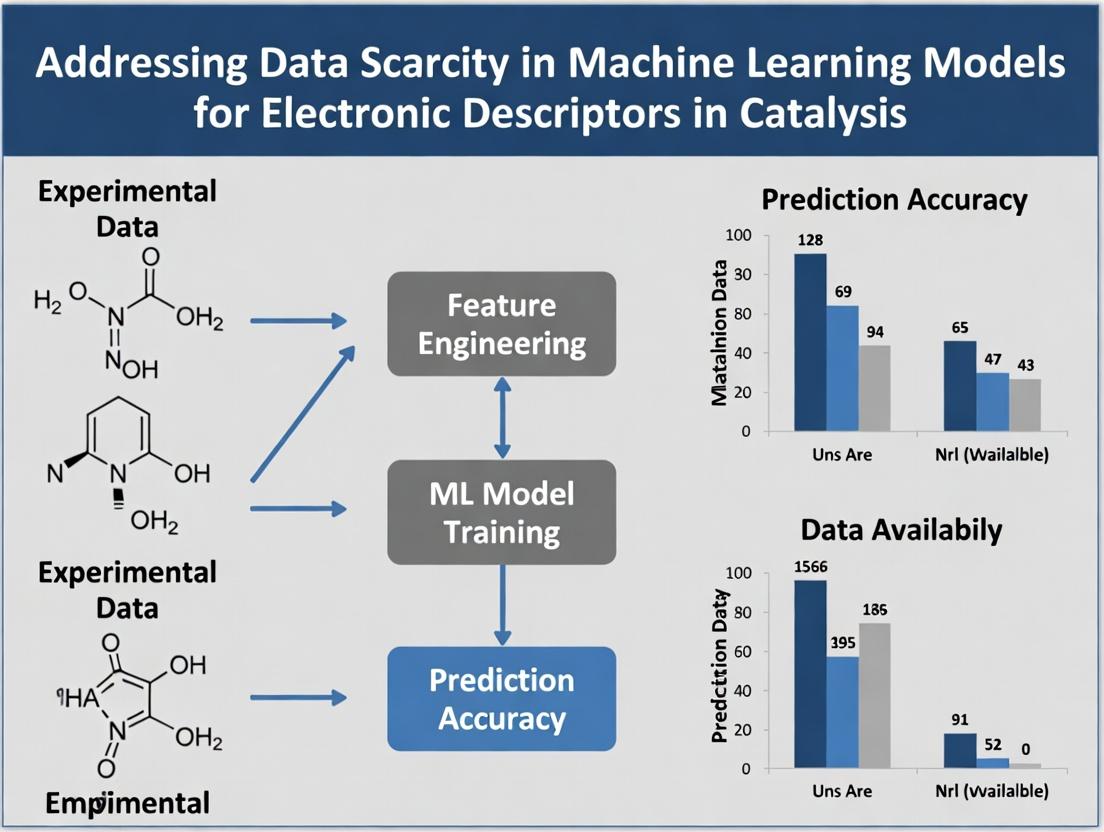
This article provides a comprehensive guide for researchers and drug development professionals facing the critical challenge of limited labeled data in electronic descriptor-based machine learning (ML) for molecular property prediction.
Beyond the Data Desert: Advanced Strategies to Overcome Data Scarcity in Molecular Property Prediction
Abstract
This article provides a comprehensive guide for researchers and drug development professionals facing the critical challenge of limited labeled data in electronic descriptor-based machine learning (ML) for molecular property prediction. We explore the fundamental causes and impacts of data scarcity, then delve into practical methodological solutions including data augmentation, transfer learning, and active learning. The guide further addresses common pitfalls and optimization strategies for model robustness, and concludes with rigorous validation frameworks and comparative analyses of emerging techniques. Our synthesis aims to equip practitioners with the knowledge to build more reliable and generalizable predictive models, accelerating discovery in computational chemistry and drug development.
The Data Scarcity Challenge: Why Small Datasets Stunt AI-Driven Molecular Discovery
Technical Support & Troubleshooting Center
FAQs
Q1: Our cell-based assay for compound screening is yielding inconsistent viability readouts, increasing cost per data point. What are the primary troubleshooting steps?
A: Inconsistent viability data often stems from cell culture health or assay protocol drift. Follow this systematic check:
- Passage Number & Contamination: Check mycoplasma contamination via PCR. Use cells below passage 20.
- Seeding Density Optimization: Re-optimize density for your plate format using a positive control. See Table 1 for common errors.
- Edge Effect Mitigation: Use a humidified chamber, pre-warm media, and utilize perimeter columns for buffer only. Consider specialized microplates.
- Compound Solvent Matching: Ensure the DMSO (or other solvent) concentration is identical (typically ≤0.5%) across all wells, including controls.
Q2: Our SPR (Surface Plasmon Resonance) runs show high non-specific binding, wasting expensive protein and ligand. How can we improve surface chemistry?
A: High background binding compromises data quality and throughput. Address it as follows:
- Surface Regeneration Scouting: Perform a regeneration scouting experiment using a pH gradient (e.g., Glycine 1.5-3.0) to find optimal conditions without damaging the chip.
- Reference Surface & Blocking: Always use a dedicated reference flow cell. Implement a blocking step with an inert protein (e.g., 0.1% BSA) or carboxymethyl dextran blockers after immobilization.
- Running Buffer Optimization: Increase ionic strength (e.g., 150-500 mM NaCl), add a mild detergent (0.005% P20), or include a chelating agent (EDTA) if applicable.
Q3: HPLC purification for compound libraries is a bottleneck. How can we increase throughput without compromising purity for ML model training?
A: To scale purification, consider these protocol modifications:
- Switch to UPC2/SFC: For chiral or normal-phase separations, Ultra-Performance Convergence Chromatography (SFC) offers faster run times and lower solvent consumption than HPLC.
- Implement Gradient Screening: Use a short, scouting gradient (e.g., 5-100% organic modifier over 1 min on a narrow column) to quickly determine optimal conditions before scaling.
- Automated Fraction Triggering: Use MS-triggered fraction collection to increase accuracy and reduce manual collection time, ensuring high-purity samples for model training.
Q4: Our biochemical assay data shows high Z' factor variability, leading to unreliable hit identification. What are key optimization parameters?
A: A Z' factor < 0.5 indicates an unreliable assay. Key optimization targets include:
- Enzyme Stability: Aliquot and freeze enzyme stocks; use a fresh aliquot daily. Include a stability time course.
- Substrate QC: Verify substrate concentration spectrophotometrically. Ensure it is at saturation (Km).
- Signal Dynamic Range: Titrate both enzyme and substrate to maximize the signal-to-background ratio. See Table 2 for target parameters.
Experimental Protocols
Protocol 1: High-Throughput qPCR for Gene Expression Validation (96-well format) Objective: Generate reproducible, quantitative gene expression data for ML model training on compound mechanism.
- Cell Lysis & Reverse Transcription: Plate cells in 96-well culture plate. Treat with compounds. Lyse cells directly with 20 µL of TRIzol/well. Perform cDNA synthesis using a high-efficiency reverse transcriptase master mix (e.g., SuperScript IV) in a total volume of 10 µL.
- qPCR Setup: Dilute cDNA 1:5 in nuclease-free water. Prepare qPCR master mix containing SYBR Green dye, forward/reverse primers (200 nM final), and 2 µL diluted cDNA per 10 µL reaction. Run in technical triplicates.
- Data Analysis: Calculate ∆∆Ct values using housekeeping gene (GAPDH) and DMSO vehicle control. Export fold-change values for model ingestion.
Protocol 2: Immobilization of His-Tagged Protein on SPR Chip (Series S Sensor Chip NTA) Objective: Generate a stable, active protein surface for kinetic binding assays.
- Chip Priming: Dock the chip and prime the system with running buffer (e.g., HBS-EP+: 10 mM HEPES, 150 mM NaCl, 3 mM EDTA, 0.05% surfactant P20, pH 7.4).
- NTA Activation: Inject a 1:1 mixture of 0.4 M EDC and 0.1 M NHS for 420 seconds at 10 µL/min. Inject 0.5 mM NiCl2 for 300 seconds.
- Protein Capture: Dilute His-tagged protein to 10 µg/mL in running buffer. Inject for 300-600 seconds to achieve ~50-100 Response Units (RU) capture.
- Surface Blocking: Inject 350 mM EDTA for 60 seconds to remove loosely bound nickel, then inject 0.1% BSA for 120 seconds to block non-specific sites.
Data Presentation
Table 1: Common Cell-Based Assay Errors & Impact on Cost
| Error Source | Typical Consequence | Estimated Cost Impact (Per 384-well Plate) | Mitigation Strategy |
|---|---|---|---|
| Inconsistent Cell Seeding | High CV (>20%), plate failure | $500 (reagents + labor) | Use automated liquid handler, validate count |
| Edge Evaporation ("Edge Effect") | False positives/negatives in outer wells | $250 (lost data points) | Use plate sealers, humidity chambers |
| Compound Precipitation | Non-linear dose response, artifact | $300 (compound wasted) | Pre-filter compounds, use DMSO gradient |
| Contaminated Cell Stock | Uninterpretable results, project delay | $1000+ (full repeat) | Regular mycoplasma testing, use low-passage aliquots |
Table 2: Target Parameters for Robust Biochemical Assay Development
| Parameter | Optimal Value | Acceptable Range | Method for Measurement |
|---|---|---|---|
| Z' Factor | > 0.7 | 0.5 - 1.0 | (1 - (3σhigh + 3σlow)/|µhigh - µlow|) |
| Signal-to-Background (S/B) | > 10 | > 3 | Mean signal of high control / mean of low control |
| Coefficient of Variation (CV) | < 10% | < 15% | (Standard Deviation / Mean) * 100 |
| Assay Window | > 10-fold | > 3-fold | Dynamic range between high and low controls |
Visualizations
Diagram 1: Data Scarcity Impact on ML Model Pipeline
Diagram 2: SPR Assay Troubleshooting Workflow
The Scientist's Toolkit: Research Reagent Solutions
| Item/Category | Example Product/Brand | Primary Function in Context |
|---|---|---|
| Automated Liquid Handler | Beckman Coulter Biomek, Integra Assist Plus | Enables precise, high-throughput cell seeding and compound transfer, reducing plate-to-plate variability. |
| Specialized Microplates | Corning Spheroid, Greiner µClear | Minimizes edge effects, enhances imaging, or supports 3D cell culture for more physiologically relevant data. |
| Ready-to-Assay Kits | Eurofins DiscoverX KINOMEscan, Thermo Fisher Z'-LYTE | Provides highly validated, off-the-shelf biochemical assays, lowering initial optimization cost and time. |
| SPR Chip & Reagents | Cytiva Series S Sensor Chip NTA, GE Healthcare | Enables label-free, kinetic binding studies for protein-ligand interactions, crucial for affinity data. |
| QC'd Chemical Libraries | Selleckchem L1200, Enamine REAL | Provides large, purity-verified (>90%) compound collections for screening, ensuring data artifacts aren't from impurities. |
| Cloud Data Platform | CDD Vault, Benchling | Centralizes experimental data with metadata, facilitating clean, structured data export for ML model training. |
Troubleshooting & FAQ Center
Q1: Our ML model for predicting molecular properties performs excellently on the training/validation set but fails on new, external test compounds. What's the primary cause and how can we diagnose it? A: This is a classic symptom of overfitting due to the data bottleneck. The model has memorized noise or specific artifacts in your limited dataset rather than learning generalizable relationships between descriptors and the target property. To diagnose:
- Perform a learning curve analysis. Train multiple models on incrementally larger subsets of your data and plot performance against training set size. If performance plateaus well before using all data, a data bottleneck is likely.
- Conduct external validation. Test the model on a truly external set from a different source or time period. A significant drop (>20% in RMSE or >30% in classification metrics) indicates poor generalization.
- Use model explainability tools (e.g., SHAP) on both training and failed predictions. If the model relies on different, non-intuitive descriptors for its external predictions, it has likely learned spurious correlations.
Q2: What are the most effective techniques to mitigate overfitting when we cannot acquire more experimental data for our electronic descriptor model? A: Implement a multi-pronged strategy focused on data efficiency and model constraint:
| Technique Category | Specific Method | Implementation Note | Expected Outcome |
|---|---|---|---|
| Data Augmentation | SMILES Enumeration, 3D Conformer Generation, Adversarial Noise Injection | For electronic descriptors, adding Gaussian noise (σ=0.01-0.05) to DFT-calculated values can simulate calculation variance. | Increases effective dataset size by 5-20x, improves robustness. |
| Transfer Learning | Pre-training on large public datasets (e.g., QM9, PubChemQC) followed by fine-tuning on your small dataset. | Freeze initial layers of the network during fine-tuning. | Can reduce required task-specific data by orders of magnitude. |
| Model Regularization | Increased Dropout (rate=0.5-0.7), Weight Decay (L2 penalty), Early Stopping with strict patience. | Monitor loss on a held-out validation set not used for training. | Reduces model complexity, forcing it to learn more robust features. |
| Simpler Architectures | Switch from deep neural networks to Gradient Boosting Machines (GBM) or Ridge Regression when N < 10,000. | GBM with ≤100 trees often outperforms DNN on small, structured descriptor data. | Lower model capacity reduces overfitting risk. |
Q3: How do we reliably estimate model performance and uncertainty when working with a small dataset (<500 samples)? A: Traditional train/test splits are unreliable. Use rigorous resampling techniques:
- Nested Cross-Validation: Provides an almost unbiased performance estimate.
- Inner Loop: Optimize hyperparameters (e.g., via grid search).
- Outer Loop: Evaluate model performance with optimized parameters.
- Bootstrapping: Repeatedly sample from your dataset with replacement to create many "pseudo-datasets." Train a model on each and aggregate predictions. This yields:
- A robust performance estimate (mean across bootstrap samples).
- Prediction Intervals: Calculate the standard deviation of the bootstrap predictions for each sample to quantify uncertainty. High uncertainty highlights areas where the model is extrapolating due to data scarcity.
Detailed Protocol: Nested Cross-Validation for Small Data
- Define Outer K-folds: Split your entire dataset into K folds (e.g., K=5 or 10).
- Iterate Outer Loop: For each outer fold i: a. Hold out fold i as the test set. b. The remaining K-1 folds form the development set. c. Inner Loop: Perform a separate K-fold cross-validation only on the development set to select the best hyperparameters (e.g., learning rate, hidden layer size). d. Train a final model on the entire development set using the best hyperparameters. e. Evaluate this final model on the held-out outer test set (fold i). Record the metric (e.g., R², MAE).
- Aggregate: The average performance across all K outer test folds is your final performance estimate. The standard deviation indicates its stability.
Q4: Our model's predictions are sensitive to minor variations in DFT calculation parameters (e.g., basis set, functional). How can we build a model robust to this "descriptor noise"? A: This is a data consistency issue. The model is learning precise numerical values that are not physically invariant.
- Solution: Incorporate data variance directly during training.
- Protocol: For each molecule in your training set, calculate its electronic descriptors using 2-3 different reasonable DFT parameter sets (e.g., B3LYP/6-31G, M062X/6-311+G). Do not treat these as separate samples. Instead, for each molecule, use the mean descriptor vector as the input feature, and append the standard deviation of each descriptor across the parameter sets as additional input features (or as an uncertainty weighting in the loss function). This explicitly teaches the model which descriptor dimensions are stable and which are noisy.
Key Research Reagent Solutions
| Item / Solution | Function in Addressing Data Scarcity |
|---|---|
| Pre-trained Foundation Models (e.g., ChemBERTa, GPT-3 for molecules) | Provides transferable molecular representations, reducing the need for massive labeled datasets specific to your property. |
| Automated First-Principles Calculation Suites (e.g., AutoGULP, ASE, high-throughput DFT workflows) | Enables systematic generation of consistent electronic descriptor data for augmentation or active learning loops. |
| Active Learning Platforms (e.g., ChemML, deepchem) | Algorithms that iteratively select the most informative molecules for expensive experimental or computational characterization, maximizing data efficiency. |
| Uncertainty Quantification Libraries (e.g., GPyTorch for Gaussian Processes, Deep Ensembles) | Provides tools to implement models that output both a prediction and its confidence, critical for reliable deployment with small data. |
| Standardized Benchmark Datasets (e.g., MoleculeNet, OCELOT) | Provides curated, high-quality data for pre-training and reliable performance comparison against state-of-the-art methods. |
Visualization: Experimental Workflow for Robust Small-Data ML
Diagram Title: Small-Data ML Workflow for Robust Models
Visualization: The Bottleneck Effect on Model Generalization
Diagram Title: Data Bottleneck Effect on Model Outcomes
Technical Support Center
Troubleshooting Guides & FAQs
Q1: My ML model for predicting HOMO/LUMO levels from SMILES strings is underperforming. What could be the source of data scarcity and how can I troubleshoot it? A: This is a core symptom of data scarcity in electronic descriptor research. First, validate your dataset's scope.
- Troubleshooting Steps:
- Audit Data Source: Check if your training data originates from a single computational method (e.g., only DFT at B3LYP/6-31G*). Models trained on single-source data fail to generalize.
- Check Property Range: Calculate the range and distribution of your target HOMO/LUMO values. Gaps or extreme clustering in property space indicate inadequate coverage.
- Verify Structural Diversity: Perform a similarity analysis (e.g., Tanimoto fingerprints) on your molecular set. High average similarity (>0.7) suggests a lack of diverse scaffolds.
- Solution: Augment data with calculations from different levels of theory (e.g., HF, ωB97X-D) or high-throughput experimentation (e.g., cyclic voltammetry screening). Use active learning to target calculations for molecules filling gaps in chemical/ property space.
Q2: When generating charge-transfer descriptors, my quantum calculations fail to converge for large, flexible drug-like molecules. How do I proceed? A: Convergence failures are common and limit data generation.
- Troubleshooting Protocol:
- Geometry Pre-optimization: Use a faster, semi-empirical method (e.g., GFN2-xTB) to generate a reasonable starting geometry before initiating higher-level DFT calculations.
- Basis Set & Functional Adjustment: Start with a smaller basis set (e.g., 6-31G*) and a robust functional (e.g., PBE), then refine with larger basis sets.
- Solvent Model: If using an implicit solvent model, try running the initial optimization in vacuo first, then add the solvent model for the final single-point energy calculation.
- Solution: Implement a tiered computational workflow that starts with inexpensive methods and only escalates computationally intensive steps for molecules that pass initial convergence checks. Document all failures as they inform the boundaries of your dataset.
Q3: The experimental electrochemical band gap I measured differs significantly from the DFT-calculated HOMO-LUMO gap. How do I reconcile this for model training? A: This discrepancy is a key data alignment challenge.
- Diagnostic Guide:
- Understand the Fundamentals: The DFT Kohn-Sham HOMO-LUMO gap is not a quasiparticle band gap. It typically underestimates the experimental optical/electrochemical gap. A systematic offset is expected.
- Calibrate Your Calculation: Establish a linear correlation between calculated gaps and experimental gaps for a small set of reference compounds within your chemical class.
- Check Experimental Conditions: Verify that your experimental gap (from CV) is corrected for the reference electrode and solvent effects. Use a consistent internal standard (e.g., ferrocene/ferrocenium).
- Solution: Do not mix raw calculated and experimental gaps directly. Use the calculated gap as a descriptor and the experimental value as the target. Alternatively, apply a calibrated scaling factor (from step 2) to the calculated values before use, clearly documenting this transformation.
Q4: I lack experimental data for excited-state descriptors (e.g., triplet energy T1). How can I create a reliable dataset for photoredox catalyst screening? A: This highlights the scarcity of high-quality excited-state data.
- Methodology for Data Generation:
- High-Throughput Computational Protocol: Use Time-Dependent DFT (TD-DFT) with a functional known for reasonable accuracy for excited states (e.g., ωB97X-D, CAM-B3LYP) and a moderate basis set (e.g., def2-SVP). Automate the workflow for thousands of candidates.
- Data Curation Critical Step: Manually check results for a random subset. Validate by comparing against any available experimental data for known catalysts (see table below). Flag and reinvestigate outliers.
- Uncertainty Quantification: Record the energy difference between the first singlet (S1) and triplet (T1) states. A large S1-T1 gap may indicate poor TD-DFT performance for that molecule.
- Solution: Create a benchmark dataset of computed descriptors for a diverse virtual library. Clearly label the computational method and its known limitations. This structured, large-scale computed dataset is valuable despite the lack of experiment.
Q5: My descriptor-based virtual screening identified hits, but they failed in subsequent assays. Could missing descriptors be the cause? A: Yes, this often points to "descriptor blindness" – your feature set lacks critical information.
- Root Cause Analysis:
- Gap Analysis: Compare your descriptor set against a comprehensive list (e.g., Mordred, DRAGON). Are you missing key classes like topological charge indices, 3D-MoRSE descriptors, or wavelet coefficients?
- Failure Mode Correlation: Analyze if the assay failures share a common chemical substructure not captured by your 2D fingerprints. This may require 3D/conformer-dependent descriptors.
- Solvent & Dynamics: Your static, in-vacuo quantum descriptors may miss critical solvent interaction or molecular flexibility effects relevant to the assay.
- Solution: Enrich your descriptor set with targeted features. For solvation effects, add computed logP or explicit solvent interaction energies. For flexibility, include descriptors from multiple low-energy conformers.
Summarized Quantitative Data
Table 1: Comparison of Computational Methods for Key Electronic Descriptors
| Descriptor | Recommended Method (Balance) | High-Accuracy Method (Costly) | Typical Error vs. Experiment | Common Data Gap |
|---|---|---|---|---|
| HOMO/LUMO (eV) | DFT, B3LYP/6-31G* | GW Approximation | ±0.2-0.5 eV | Experimental electrochemical potentials for diverse, complex molecules. |
| Dipole Moment (D) | DFT, PBE0/def2-SVP | CCSD(T)/aug-cc-pVTZ | ±0.2-0.5 D | Measured moments in relevant solvent environments for drug-like compounds. |
| Polarizability (a.u.) | HF/6-31+G* | DFT, ωB97X-D/aug-cc-pVTZ | ±2-5% | Experimental values for large, conjugated systems beyond benchmark sets. |
| Triplet Energy T1 (eV) | TD-DFT, ωB97X-D/def2-TZVP | CASPT2 | ±0.3-0.6 eV | Systematic experimental T1 data from phosphorescence for organic molecules. |
| Fukui Indices | DFT, B3LYP/6-31G* (N+1, N-1) | Finite Difference Cond. | Qualitative | Experimental validation via kinetic or spectroscopic probes of site reactivity. |
Table 2: Public Data Repository Coverage Analysis
| Repository | Primary Content | Estimated Compounds with Electronic Descriptors | Key Limitation (Source of Scarcity) |
|---|---|---|---|
| QM9 | Small organic molecules (≤9 heavy atoms) | 134k (Geometry, Energy, Props) | Size/scope irrelevant to drug discovery; no excited states. |
| Harvard CEP | Organic photovoltaic candidates | ~3.3M (DFT HOMO/LUMO) | Single level of theory (PBE0); limited experimental validation. |
| PubChemQC | DFT calculations for PubChem | ~4.2M (at B3LYP/6-31G*) | Homogeneous method; contains failures; no post-HF corrections. |
| NOMAD | Diverse computational materials science | ~100M entries (varies widely) | Heterogeneous data format and quality; difficult to curate. |
| Experimental: OCELOT | Curated experimental optoelectronic data | ~1.2k (from literature) | Extremely small scale relative to chemical space; curation bottleneck. |
Experimental Protocols
Protocol 1: Generating a Benchmark Dataset for HOMO/LUMO Using Combined DFT & Experiment Objective: Create a high-quality, aligned dataset for ML training.
- Compound Selection: Curate a diverse set of 500-1000 organic molecules with available commercial sourcing or synthesis pathways.
- Computational Descriptor Generation:
- Software: Use Gaussian, ORCA, or PSI4.
- Geometry Optimization: Employ DFT/B3LYP with 6-31G* basis set in vacuo. Confirm no imaginary frequencies.
- Single Point Energy: Re-calculate at a higher level (e.g., ωB97X-D/def2-TZVP) with an implicit solvent model (e.g., IEFPCM for acetonitrile).
- Extract: HOMO, LUMO, Dipole Moment, Mulliken Electronegativity.
- Experimental Validation:
- Cyclic Voltammetry (CV): Prepare 1 mM solution of each compound in dry acetonitrile with 0.1 M TBAPF6 as supporting electrolyte. Use Ag/Ag+ reference electrode and standard ferrocene/ferrocenium (Fc/Fc+) internal standard.
- Measurement: Scan at 100 mV/s. Determine oxidation (Eox) and reduction (Ered) onsets.
- Alignment: Convert to HOMO/LUMO estimates: HOMO ≈ - (Eox vs. Fc/Fc+ + 4.8) eV; LUMO ≈ - (Ered vs. Fc/Fc+ + 4.8) eV.
- Data Curation: Tabulate computed and experimental values. Flag compounds with large discrepancies (>0.4 eV) for methodological review.
Protocol 2: High-Throughput Workflow for Excited-State Descriptors Objective: Calculate triplet energy (T1) and spin-density descriptors for a virtual library.
- Input Preparation: Generate a SMILES list. Use RDKit to generate an initial 3D conformation for each.
- Automated Quantum Chemistry Pipeline (e.g., using AQME):
- Step 1 - Pre-optimization: Apply GFN2-xTB to refine geometry.
- Step 2 - DFT Optimization: Use r^2SCAN-3c/def2-mTZVP for robust ground-state optimization.
- Step 3 - TD-DFT Calculation: Perform TD-DFT (Tamm-Dancoff approximation) at the PBE0/def2-SVP level to obtain the first 5 triplet excited states.
- Step 4 - Descriptor Extraction: Parse output files for: T1 energy (eV), S1-T1 gap, excited-state dipole moment, and spin density distribution (cube files).
- Quality Control: Scripts automatically check for convergence, negative excitations, and unrealistic energies. Calculations that fail are rerouted with modified parameters (e.g., increased SCF cycles).
- Database Storage: Store results in a structured SQL/NoSQL database with metadata (SMILES, charge, multiplicity, functional, basis set, convergence status).
Visualizations
Title: Computational Descriptor Generation Workflow
Title: Electronic Descriptor Data Gaps & ML Impact
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Resources for Electronic Descriptor Research
| Item / Solution | Function / Purpose | Example / Specification |
|---|---|---|
| Quantum Chemistry Software | Perform electronic structure calculations to generate descriptors from first principles. | ORCA (Free, powerful), Gaussian/GaussView (Industry standard), Psi4 (Open-source). |
| Cheminformatics Library | Handle molecular I/O, generate fingerprints, calculate simple molecular descriptors. | RDKit (Open-source, Python/C++), Open Babel (File conversion). |
| High-Performance Computing (HPC) Access | Essential for large-scale quantum calculations on thousands of molecules. | Local cluster, Cloud HPC (AWS, Azure), National supercomputing centers. |
| Benchmark Experimental Kit | Validate computed descriptors with gold-standard measurements. | Potentiostat for CV, UV-Vis-NIR Spectrometer for optical gaps, Glovebox (for air-sensitive electrochemistry). |
| Descriptor Calculation Software | Generate comprehensive descriptor sets beyond basic QM outputs. | Dragon (Commercial, ~5000 descriptors), Mordred (Open-source RDKit wrapper, ~1800 descriptors). |
| Curated Experimental Database | Source for validation data and co-training ML models. | OCELOT (Optoelectronic), NIST Computational Chemistry Comparison (CCCBDB). |
| Automation & Workflow Tool | Manage, automate, and reproduce computational pipelines. | AiiDA (Materials science), AQME (Automated QM workflows), Snakemake/Nextflow (General workflow managers). |
| ML Framework with Graph Support | Train models directly on molecular graphs or complex descriptor sets. | PyTorch Geometric, DGL-LifeSci, scikit-learn (for tabular data). |
Troubleshooting Guide & FAQs
Q1: My model for predicting acute oral toxicity shows excellent validation accuracy but fails dramatically on new, structurally distinct compounds. What could be the issue?
A1: This is a classic sign of dataset bias and overfitting in low-data regimes. The training data likely lacks chemical diversity, causing the model to learn narrow, non-generalizable patterns.
- Troubleshooting Steps:
- Analyze Applicability Domain: Use distance-based (e.g., leverage, Euclidean) or similarity-based (Tanimoto) metrics to quantify how different your new compounds are from the training set.
- Employ Ensemble Methods: Combine predictions from models trained on different descriptor sets (e.g., ECFP fingerprints, Mordred descriptors, and QM properties) to increase robustness.
- Implement Data Augmentation: Use SMILES enumeration or realistic (non-arbitrary) molecular atom/bond masking to artificially expand your limited training set.
Q2: When predicting solubility, my ML model performs poorly on zwitterionic compounds despite good overall performance. How can I address this specific blind spot?
A2: The model's descriptors likely fail to capture the complex, pH-dependent ionization state crucial for zwitterion solubility.
- Troubleshooting Steps:
- Incorporate State-Specific Descriptors: Calculate and use descriptors for the major microspecies at the target pH (e.g., using
RDKitorOpenBabel). Key descriptors should include partial charges, dipole moment, and hydrogen bond donor/acceptor counts for the correct ionization form. - Use Transfer Learning: Pre-train a model on a larger, general solubility dataset (e.g., AqSolDB), then fine-tune the last layers using your smaller, specialized dataset that includes zwitterions.
- Adopt a Multi-Task Approach: Jointly train the model to predict both solubility and a related property like pKa, which forces the model to learn underlying ionization physics.
- Incorporate State-Specific Descriptors: Calculate and use descriptors for the major microspecies at the target pH (e.g., using
Q3: In binding affinity prediction, how do I handle missing 3D structural information for protein targets, which is common in low-data scenarios?
A3: Rely on ligand-based or simplified structure-based methods when full 3D complexes are unavailable.
- Troubleshooting Steps:
- Use Pharmacophore Fingerprints: Generate fingerprints that encode the spatial arrangement of key functional features, derived from a single known active ligand or a minimal set.
- Leverage Protein Sequence Descriptors: Instead of 3D structure, use protein sequence-derived features (e.g., amino acid composition, PSI-BLAST profiles, pre-trained language model embeddings) as input alongside compound descriptors.
- Apply Kinase-Kernel Similarity: For kinase targets, use a specialized similarity kernel that compares proteins based on alignment of key kinase domains, enabling affinity prediction for kinases with no co-crystal structures.
Q4: My Bayesian optimization loop for molecular design suggests compounds that are synthetically intractable. How can I constrain the generation?
A4: Integrate synthetic feasibility rules or costs directly into the objective function or search space.
- Troubleshooting Steps:
- Use a SA Score Penalty: Incorporate the Synthetic Accessibility (SA) Score from
RDKitas a penalty term in your acquisition function:Adjusted Score = Predicted Affinity - λ * SA_Score. - Employ a Reaction-Based Generator: Use a generative model (like a GVAE or GAN) that builds molecules step-by-step from available building blocks using known chemical reaction templates, ensuring tractability by construction.
- Post-Filter with Retrosynthesis Tools: Pass all suggested compounds through a retrosynthesis planner (e.g., AiZynthFinder, ASKCOS) and filter out those with no plausible synthetic pathway under user-defined constraints.
- Use a SA Score Penalty: Incorporate the Synthetic Accessibility (SA) Score from
Experimental Protocols
Protocol 1: Evaluating Model Generalizability with Cluster-Based Splitting
Objective: To create training/test splits that rigorously assess a model's ability to generalize to novel chemical scaffolds, mitigating over-optimistic random splitting.
- Descriptor Calculation: Generate extended-connectivity fingerprints (ECFP4, radius=2) for all molecules in the dataset using
RDKit. - Similarity Matrix: Compute the pairwise Tanimoto similarity matrix.
- Clustering: Apply the Butina clustering algorithm (with a similarity cutoff of 0.6) to group structurally similar molecules.
- Stratified Split: Randomly allocate entire clusters to either the training (80%) or test set (20%), ensuring no structurally similar molecules leak between sets.
- Model Training & Evaluation: Train the model on the training clusters. Evaluate its performance exclusively on the held-out clusters to measure scaffold generalization.
Protocol 2: Data Augmentation via SMILES Enumeration for Solubility Models
Objective: To artificially expand a small solubility dataset by representing each molecule in multiple, equally valid SMILES strings, encouraging the model to learn invariant molecular features.
- Canonicalization: Start with the canonical SMILES for each molecule in the original dataset.
- Enumeration: For each molecule, use
RDKit'sChem.MolToSmiles()function in a non-canonical mode to generate up to 50 random, valid SMILES representations. The exact number can be tuned based on dataset size. - Label Assignment: Assign the same measured solubility value (logS) to every SMILES string derived from the same original molecule.
- Model Input: Use these augmented SMILES strings as direct input for a sequence-based model (e.g., LSTM, Transformer) or calculate fingerprints from each variant for a descriptor-based model.
Protocol 3: Protein-Ligand Affinity Prediction using Sequence-Based Descriptors
Objective: To predict binding affinity (pKi/pIC50) for protein-ligand pairs without 3D structural data, using only protein sequences and ligand SMILES.
- Protein Feature Generation:
- Input the target protein's amino acid sequence.
- Use the
protbert-bfdpre-trained model from thetransformerslibrary to generate per-residue embeddings. - Apply global mean pooling across the sequence to obtain a fixed-length (1024-dim) protein descriptor vector.
- Ligand Feature Generation:
- Input the compound's SMILES.
- Calculate a 2048-bit ECFP4 fingerprint and a 200-dim learned embedding from a pre-trained
ChemBERTamodel. Concatenate them.
- Data Integration & Modeling:
- Concatenate the protein descriptor vector and the combined ligand feature vector.
- Feed the fused vector into a fully connected neural network (e.g., 3 hidden layers with dropout) to perform regression for the affinity value.
- Train using a mean squared error loss on a benchmark dataset like PDBbind.
Table 1: Performance of Low-Data ML Models on Toxicity Endpoints
| Model Type | Dataset (Size) | Endpoint | Metric (Value) | Key Limitation in Low-Data Context |
|---|---|---|---|---|
| Random Forest (Mordred) | EPA ToxCast (~1k cpds) | Nuclear Receptor | BA: 0.78 | High-dimensional descriptors lead to overfit |
| Graph Neural Network (GIN) | ClinTox (~1.5k cpds) | Hepatotoxicity | AUC: 0.71 | Requires careful hyperparameter tuning |
| Support Vector Machine | LD50 (~8k cpds) | Acute Oral Toxicity | Acc: 0.85 | Poor calibration on out-of-domain scaffolds |
Table 2: Solubility Prediction Methods & Data Requirements
| Method | Typical Data Requirement | Avg. RMSE (logS) | Advantage for Low-Data | Disadvantage |
|---|---|---|---|---|
| Abraham Solvation Equation | ~100s (curated) | 0.6 - 0.8 | Physicochemically interpretable | Limited to congeneric series |
| Ensemble (RF/XGB) on AqSolDB | ~10,000 | 0.7 - 1.0 | Robust, off-the-shelf | Generalizes poorly to exotic chemotypes |
| Fine-Tuned ChemBERTa | ~1,000 (specialized) | 0.5 - 0.7 | Leverages pretraining on large corpuses | Computationally intensive to fine-tune |
Table 3: Binding Affinity Prediction Without High-Resolution Structures
| Approach | Input Data | PDBbind Core Set RMSE (pK) | Ideal Low-Data Use Case |
|---|---|---|---|
| Classical QSAR | Ligand Descriptors Only | 1.8 - 2.2 | Single-target series with <100 compounds |
| Siamese Network | Protein Seq. + Ligand Fingerprint | 1.4 - 1.6 | Multiple related targets (e.g., kinase family) |
| Interaction Fingerprint (IFP) | 1 Known PDB Complex + Ligands | 1.6 - 1.9 | Scaffold hopping around a single reference structure |
Visualizations
Toxicity Model Generalizability Workflow
Solubility Prediction for Zwitterions
Sequence-Based Affinity Prediction Pipeline
The Scientist's Toolkit: Key Research Reagents & Solutions
| Item/Category | Function & Rationale for Low-Data Scenarios |
|---|---|
| RDKit | Open-source cheminformatics toolkit. Essential for generating molecular descriptors, fingerprints, and performing data augmentation (SMILES enumeration) without costly commercial software. |
| Pre-trained Models (ChemBERTa, ProtBERT) | Language models trained on vast corpora of chemical structures or protein sequences. Provide informative molecular/protein embeddings that serve as a knowledge-rich starting point for fine-tuning on small datasets. |
| AqSolDB / ChEMBL | Publicly available, curated databases for solubility and bioactivity. Serve as source data for pre-training or as external validation sets to assess model generalizability. |
| Applicability Domain Tools (e.g., ADAN) | Software/packages to calculate the applicability domain of QSAR models. Critical for identifying when low-data models are being asked to make predictions outside their reliable scope. |
| Bayesian Optimization Libraries (BoTorch, GPyOpt) | Enable efficient navigation of chemical space with minimal experiments. Crucial for optimizing molecular properties when synthesis and testing capacity (data generation) is severely limited. |
| Synthetic Accessibility Scorers (SA Score, RA Score) | Algorithms that estimate the ease of synthesizing a proposed molecule. Must be integrated into generative AI pipelines to ensure suggested compounds are practical, addressing a major failure mode in low-data design. |
Bridging the Gap: Practical Data-Centric and Model-Centric Solutions for Researchers
Troubleshooting Guides & FAQs
SMILES Enumeration
Q1: After canonicalizing enumerated SMILES strings, my dataset size reduces instead of increasing. What is the issue? A: This occurs when the canonicalization algorithm (e.g., from RDKit) maps all enumerated variants of the same molecule back to an identical canonical string. This is expected behavior, not an error. The augmentation's value is in exposing the model to diverse SMILES representations during training, not in permanently expanding the stored dataset. Implement on-the-fly enumeration within your data loader.
Q2: My model fails to learn from enumerated SMILES, showing high training loss. A: This often indicates an issue with SMILES parsing or tokenization.
- Check Validity: Use RDKit's
Chem.MolFromSmiles()on a sample of enumerated strings to ensure they generate valid molecules. Invalid SMILES can corrupt training. - Review Tokenization: Ensure your tokenizer's vocabulary includes all symbols (e.g., parentheses, ring digits, '=') generated during enumeration. Missing tokens lead to failures.
Q3: Are there best practices for choosing the number of SMILES variants per molecule? A: There is a diminishing return. Excessive enumeration can bias the dataset. A common starting point is 10-50 variants per molecule. Monitor model performance on a validation set to find the optimal point.
3D Conformer Generation
Q4: Conformer generation with RDKit's ETKDG is extremely slow for my dataset of >10k molecules. How can I speed it up? A: The ETKDG algorithm is computationally intensive. Consider these steps:
- Parallelize: Use Python's
multiprocessinglibrary to distribute conformer generation across CPU cores. - Optimize Parameters: Reduce
numConfs(the number of conformers to generate per molecule) for the initial augmentation pass. You can generate fewer, more diverse conformers usingprunermsThreshandclusterRMSThreshparameters. - Pre-filter: Use a faster 2D similarity filter to select a diverse subset of molecules for 3D augmentation if full-set generation is prohibitive.
Q5: How do I handle conformer generation for molecules with undefined stereochemistry? A: RDKit may fail or produce unrealistic conformers. Implement a pre-processing step:
- Identify molecules with undefined tetrahedral centers (
ChiralType.CHI_UNSPECIFIED). - For each, enumerate possible stereoisomers using
EnumerateStereoisomers(). - Generate conformers for each defined stereoisomer separately.
- Select the lowest energy conformer from all stereoisomers or use all for augmentation.
Q6: What metrics should I use to ensure the quality of generated conformers? A: Common quality checks include:
- Energy Strain: Compare MMFF94 or UFF energy of the generated conformer to a minimized version.
- Steric Clash: Check for unrealistic atom-atom distances (van der Waals overlaps).
- Experimental Comparison: If crystal structures are available, calculate Root-Mean-Square Deviation (RMSD) of atomic positions.
Table 1: Performance Comparison of Conformer Generation Methods (Approximate Timings)
| Method | Software/Tool | Speed (mols/sec)* | Handling of Uncertainty | Recommended Use Case |
|---|---|---|---|---|
| ETKDGv3 | RDKit | ~1-5 | Requires defined stereochemistry | Standard small organic molecules. |
| OMEGA | OpenEye | ~10-50 | Excellent stereoisomer handling | Production-scale, high-quality conformers. |
| ConfGenx | Schrödinger | ~5-20 | Robust force field | Drug-like molecules in lead optimization. |
| Distance Geometry (Basic) | RDKit (basic) | ~10-20 | Poor | Fast, low-quality baseline. |
*Speed is hardware-dependent and estimated for typical drug-like molecules.
Noise Injection
Q7: When adding Gaussian noise to atomic coordinates or electronic descriptors, what is a principled way to set the noise level (σ)? A: The noise level should be relative to the natural variation in your data.
- For Atomic Coordinates: Calculate σ as a fraction (e.g., 0.01-0.05) of the average bond length in your dataset (~1.5 Å). Start with σ = 0.05 Å.
- For Electronic Descriptors: Calculate the standard deviation of each descriptor column. Set σ for each descriptor to 0.01-0.1 times its standard deviation. Always validate that noise does not create physically impossible values (e.g., negative energies).
Q8: Noise injection causes some molecular graphs to become invalid (e.g., broken bonds, extreme angles). How should this be handled? A: You must implement a validity check and a rejection/repair strategy.
- Strategy 1 (Rejection): After injecting noise, compute inter-atomic distances. If any bonded atom pair exceeds a threshold (e.g., 2x typical bond length), discard that augmented sample.
- Strategy 2 (Repair): Apply a mild force field minimization (e.g., 50 steps of UFF in RDKit) to "relax" the noisy structure back to a physically plausible state.
Q9: Can noise be applied to the molecular graph adjacency matrix? A: Yes, but with caution. Randomly adding/removing edges (bonds) can drastically alter chemistry. A safer graph-level augmentation is atom/ bond masking, where a small fraction of node or edge features are randomly set to zero, forcing the model to use context for prediction.
Experimental Protocols
Protocol 1: SMILES Enumeration & Training Data Loader
Objective: Integrate stochastic SMILES augmentation into a PyTorch or TensorFlow training pipeline.
- Store only the canonical SMILES for each molecule in your dataset.
- In your
Datasetclass's__getitem__method: a. Retrieve the canonical SMILES string for the given index. b. Use RDKit to create a molecule object (Chem.MolFromSmiles). c. Generate a random, non-canonical SMILES string for the molecule (Chem.MolToSmiles(mol, doRandom=True, canonical=False)). d. Tokenize this random SMILES string using your predefined tokenizer. e. Return the tokenized sequence and the associated label (e.g., property value). - This ensures the model sees a different textual representation of each molecule in every epoch.
Protocol 2: High-Quality 3D Conformer Generation with ETKDG
Objective: Generate a diverse, low-energy set of conformers for a molecule with defined stereochemistry.
- Input: A molecule (
mol) with sanitized chemistry and defined stereocenters. - Setup Parameters:
- Generate:
conf_ids = AllChem.EmbedMultipleConfs(mol, params=params) - Minimize & Score: For each conformer ID, perform a short UFF minimization (
AllChem.UFFOptimizeMolecule) and calculate its energy (AllChem.UFFGetMoleculeForceField). - Cluster & Select: Cluster conformers based on heavy-atom RMSD (e.g., threshold=1.0 Å). Select the lowest-energy conformer from each cluster for the final augmented set.
Protocol 3: Noise Injection on Quantum Mechanical Descriptors
Objective: Create augmented samples for electronic property prediction models.
- Input: A matrix
Xof size (nmolecules, ndescriptors). Each column is a descriptor (e.g., HOMO, LUMO, dipole moment). - Calculate Statistics: Compute the standard deviation
std_devfor each descriptor column across the training set only. - Define Noise Scale: Set a global scaling factor
alpha(e.g., 0.05). - Generate Augmented Batch: For a batch of data
X_batch, generate a noise matrixNof the same shape, where each element is sampled fromNormal(0, 1). The augmented batch is:X_augmented = X_batch + alpha * N * std_dev(broadcasted). - Clip (Optional): Clip values to a physically plausible range (e.g., HOMO energy cannot be >0).
Diagrams
Diagram 1: Molecular Data Augmentation Workflow
Diagram 2: Thesis Context in Addressing Data Scarcity
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Tools & Libraries for Molecular Augmentation Experiments
| Item / Software | Primary Function in Augmentation | Key Considerations / Notes |
|---|---|---|
| RDKit (Open Source) | Core toolkit for SMILES manipulation, 2D/3D molecular operations, and ETKDG conformer generation. | The fundamental library. Use rdkit.Chem and rdkit.Chem.AllChem. |
| OpenEye Toolkit (Commercial) | Industry-standard for high-speed, high-quality conformer generation (OMEGA) and molecular modeling. | Superior handling of stereochemistry and conformational sampling; requires license. |
| PyTorch / TensorFlow | Deep learning frameworks for building models and implementing custom data augmentation layers/data loaders. | Essential for integrating on-the-fly augmentation into the training pipeline. |
| PyMOL / VMD | Molecular visualization software. | Critical for qualitatively validating the 3D structures of generated conformers. |
| Good-Turing Frequency Estimator | Statistical method to assess the coverage of chemical space by your augmented dataset. | Helps answer "Has augmentation introduced meaningful new information?" |
| UFF/MMFF94 Force Fields (in RDKit) | Used to minimize and score generated 3D conformers, ensuring physical realism. | Apply after noise injection or conformer generation to "clean" structures. |
| scikit-learn | Used for preprocessing descriptors (e.g., StandardScaler) and calculating statistics for noise injection parameters. | Simple, reliable utilities for feature-space augmentation steps. |
Technical Support Center
Troubleshooting Guides & FAQs
Q1: I am fine-tuning a pre-trained model (e.g., ChemBERTa) on my small dataset of electronic descriptors, but the validation loss is not decreasing. What could be wrong? A: This is a classic symptom of overfitting or inappropriate learning rate settings.
- Check 1: Learning Rate. Pre-trained models require very low learning rates for fine-tuning. Try a range from 1e-5 to 5e-4. Use a learning rate scheduler.
- Check 2: Data Representation. Ensure your input SMILES strings or molecular graphs are tokenized/processed identically to how the base model was trained. A mismatch in tokenization will prevent effective transfer.
- Check 3: Early Stopping. Implement early stopping with a patience of 5-10 epochs to halt training when validation loss plateaus.
- Protocol: Perform a learning rate sweep. Train your model for 10 epochs using learning rates [1e-4, 5e-5, 1e-5]. Plot the validation loss to identify the optimal starting point.
Q2: How do I choose which layers of a pre-trained graph neural network (GNN) to freeze versus fine-tune for my descriptor prediction task? A: The optimal strategy depends on dataset similarity and size.
- Strategy for Very Small Data (<1k samples): Freeze all but the final prediction head (last 1-2 layers). This treats the pre-trained model as a fixed feature extractor.
- Strategy for Moderately Similar Data (1k-10k samples): Unfreeze and fine-tune the last 2-3 message-passing layers of the GNN, along with the prediction head. The early layers capture fundamental chemistry.
- Strategy for Dissimilar Data or Larger Data (>10k samples): Consider fine-tuning all layers with a very low initial learning rate, progressively unlocking layers if needed.
- Protocol:
- Start with a fully frozen backbone.
- Unfreeze the final GNN layer and the prediction head. Train for 20 epochs.
- If performance is suboptimal, unfreeze the preceding GNN layer and continue training with a reduced learning rate (by a factor of 2-5).
- Monitor validation metrics closely to avoid overfitting.
Q3: When using a model pre-trained on PubChem (e.g., 10M compounds) for a specific therapeutic area (e.g., kinase inhibitors), should I use the entire pre-trained model or just the embeddings? A: For addressing data scarcity in descriptor prediction, using the entire model with fine-tuning is generally superior. The embeddings alone discard learned complex feature interactions.
- Recommendation: Use the full model architecture. Replace the final output layer to predict your continuous electronic descriptors (e.g., HOMO/LUMO, polarizability) instead of the original pre-training task (e.g., molecular property classification).
- Protocol:
- Load the pre-trained weights (e.g., for a model like
ChemBERTaorPretrained GNN). - Remove the original classification/regression head.
- Append a new regression head suited to your output dimensionality (e.g., a 2-layer MLP for predicting HOMO and LUMO energies).
- Proceed with fine-tuning as described in Q1 & Q2.
- Load the pre-trained weights (e.g., for a model like
Q4: I encounter "CUDA out of memory" errors when fine-tuning large models. How can I proceed? A: This is a hardware limitation common with large GNNs or Transformers.
- Solution 1: Gradient Accumulation. Simulate a larger batch size by accumulating gradients over several smaller batches before performing an optimizer step.
- Solution 2: Mixed Precision Training. Use Automatic Mixed Precision (AMP) to reduce memory footprint and speed up computation.
- Solution 3: Reduce Model Footprint. Try a smaller pre-trained variant (e.g.,
ChemBERTa-uncasedvs.ChemBERTa-cased) or use gradient checkpointing. - Protocol for Gradient Accumulation:
Experimental Protocols
Protocol 1: Standard Fine-Tuning Workflow for Electronic Descriptor Prediction
- Data Preparation: Curate your small dataset of molecules and their target electronic descriptors. Split into Train/Validation/Test sets (e.g., 70/15/15).
- Model Initialization: Load a pre-trained model from a large chemical library (e.g.,
ChemBERTafrom Hugging Face,Pretrained GNNfrom MoleculeNet). - Head Replacement: Replace the model's final output layer with a new, randomly initialized regression head matching your output descriptor count.
- Layer Freezing: Initially freeze all layers of the pre-trained backbone.
- Initial Training: Train only the new head for 5-10 epochs with a moderate learning rate (e.g., 1e-3) to establish a stable baseline.
- Gradual Unfreezing: Unfreeze the pre-trained model's final 1-3 layers. Lower the learning rate (e.g., 5e-5).
- Full Fine-Tuning: Train the entire model with a very low learning rate (e.g., 1e-5), using early stopping on the validation set.
- Evaluation: Report performance (MAE, RMSE) on the held-out test set.
Protocol 2: Benchmarking Transfer Learning Efficacy
- Baseline: Train a model from scratch (no pre-training) on your small dataset. Record test set performance.
- Feature Extraction: Use the frozen pre-trained model to generate molecular embeddings. Train a simple model (e.g., Random Forest, shallow MLP) on these fixed features. Record performance.
- Fine-Tuning: Perform the full fine-tuning protocol (Protocol 1). Record performance.
- Analysis: Compare the three results in a table. Successful transfer learning should show: Fine-Tuning > Feature Extraction > Baseline.
Data Presentation
Table 1: Performance Comparison of Modeling Approaches for Predicting HOMO Energies (eV) on a Small Dataset (n=500)
| Model Approach | Pre-trained Source | MAE (Test) ± Std Dev | RMSE (Test) ± Std Dev | Training Time (min) |
|---|---|---|---|---|
| MLP (From Scratch) | N/A | 0.52 ± 0.04 | 0.68 ± 0.05 | 5 |
| Random Forest on ECFP4 | N/A | 0.41 ± 0.03 | 0.55 ± 0.04 | 2 |
| GNN (From Scratch) | N/A | 0.48 ± 0.06 | 0.65 ± 0.07 | 25 |
| ChemBERTa (Feature Extract) | PubChem 10M | 0.35 ± 0.02 | 0.48 ± 0.03 | 8 |
| ChemBERTa (Fine-Tuned) | PubChem 10M | 0.22 ± 0.01 | 0.31 ± 0.02 | 35 |
| Pretrained GNN (Fine-Tuned) | PCQM4Mv2 | 0.19 ± 0.02 | 0.28 ± 0.03 | 40 |
Visualizations
Title: Transfer Learning Workflow from Big Data to Small Data
Title: Stepwise Fine-Tuning Protocol Logic
The Scientist's Toolkit
Table 2: Essential Research Reagent Solutions for Transfer Learning Experiments
| Item | Function & Relevance |
|---|---|
| Pre-Trained Model Weights (e.g., ChemBERTa, ChemGNN) | Foundational knowledge base from large-scale chemical libraries; the core "reagent" for transfer learning. |
| Small, Curated Target Dataset | The specific electronic descriptor data (e.g., DFT-calculated properties) for the molecules of interest. |
| Deep Learning Framework (PyTorch, TensorFlow with RDKit) | Environment for loading, modifying, and training neural network models. |
| Molecular Featurizer/Tokenizer (e.g., RDKit, SMILES Tokenizer) | Converts raw molecular structures (SMILES, SDF) into the input format required by the pre-trained model. |
| Learning Rate Scheduler (e.g., ReduceLROnPlateau, Cosine Annealing) | Critically adjusts learning rate during fine-tuning to avoid catastrophic forgetting and enable convergence. |
| Automatic Differentiation & Mixed Precision (e.g., PyTorch AMP) | Enables efficient training and mitigates GPU memory constraints when handling large models. |
Model Checkpointing Library (e.g., Hugging Face transformers, PyTorch Lightning) |
Simplifies the process of saving, loading, and managing different model versions during experimentation. |
| Chemical Validation Set | A hold-out set of molecules not used in training, essential for detecting overfitting and assessing generalizability. |
Introduction to Our Technical Support Center Welcome to the technical support hub for researchers implementing Active Learning (AL) loops to combat data scarcity in electronic descriptor-based ML models for molecular discovery. This guide provides targeted troubleshooting and FAQs to optimize your experimental design cycle.
Frequently Asked Questions (FAQs)
Q1: My acquisition function consistently selects outliers, leading to poor model generalization. What should I do? A: This is often a sign of inadequate exploration-exploitation balance.
- Troubleshooting: Re-calibrate your acquisition function. For
Upper Confidence Bound (UCB), increase thekappaparameter to encourage exploration. ForExpected Improvement (EI)orProbability of Improvement (PI), consider adding a small noise term (xi) to prevent over-exploitation of small improvements. Ensembling multiple acquisition functions can also stabilize selections.
Q2: How do I handle batch selection when lab throughput is limited, but I want to run parallel experiments? A: Implement batch-aware acquisition strategies.
- Troubleshooting: Move from greedy selection to batch modes like:
- K-Means Batch Sampling: Cluster the top N candidates from the acquisition function and select the cluster centroids for diversity.
Local Penalization: Artificially reduce the acquisition function value around each selected point in the batch to encourage spatial diversity in the feature space.- Use
q-EIorq-PIwhich are explicitly designed for parallel querying.
Q3: My initial dataset is very small (<50 data points). Which model should I start my AL loop with? A: Prioritize models with strong uncertainty quantification capabilities from small data.
- Troubleshooting: Start with a Gaussian Process (GP) model. GPs provide inherent, well-calibrated uncertainty estimates (the predictive variance), which is crucial for most acquisition functions. If molecular descriptors are high-dimensional, use a sparse GP or a model combining a neural network feature extractor with a GP head (Deep Kernel Learning) to manage computational cost.
Q4: The model's uncertainty estimates seem unreliable. How can I validate them? A: Perform calibration checks on your model's predictive distribution.
- Troubleshooting Protocol:
- Reserve a small validation set from your pool data or initial seed.
- For each validation point, have your model predict both a mean (
μ) and standard deviation (σ). - Calculate the
z-score:(y_true - μ) / σ. - Plot a histogram of the z-scores. A well-calibrated model will produce a histogram that closely resembles a standard normal distribution (mean=0, variance=1).
- If miscalibrated, consider using temperature scaling or switching to a model with better inherent uncertainty quantification.
Q5: How do I know when to stop the AL loop? A: Define stopping criteria before starting the loop. Common metrics include:
- Performance Plateau: Stop when the improvement in target property (e.g., binding affinity, solubility) over the last
Kcycles falls below a thresholdΔ. - Budget Exhaustion: Pre-define a maximum number of experiments or computational budget.
- Target Achievement: Stop when a molecule meets a specific property threshold.
Experimental Protocols
Protocol 1: Setting Up a Basic Active Learning Loop for Molecular Screening
- Objective: Iteratively identify compounds with high predicted activity from a large virtual library.
- Materials: See "Research Reagent Solutions" table.
- Method:
- Seed Data: Assemble a small, diverse set of molecules (
n=50-100) with experimentally measured target properties. - Featurization: Compute electronic descriptors (e.g., HOMO/LUMO energies, dipole moment, polarizability) for all molecules in the seed and the large unlabeled pool (
~10,000-100,000) using quantum chemistry software (e.g., DFT, semi-empirical methods). - Model Training: Train a Gaussian Process Regressor (GPR) on the seed data's descriptors and target values.
- Acquisition: Use the trained GPR to predict the mean and variance for all molecules in the unlabeled pool. Apply the
Expected Improvement (EI)acquisition function to rank them. - Selection & Experiment: Select the top 5-10 molecules from the ranked list for synthesis and experimental assay.
- Update: Add the new experimental results to the seed data.
- Loop: Repeat steps 3-6 until a stopping criterion is met.
- Seed Data: Assemble a small, diverse set of molecules (
Protocol 2: Calibrating Model Uncertainty for Reliable Query
- Objective: Ensure acquisition functions act on trustworthy uncertainty estimates.
- Method:
- Split your current labeled data into 80% training and 20% calibration sets.
- Train your model (e.g., GPR, Bayesian Neural Network) on the training set.
- Predict the mean (
μ) and standard deviation (σ) for the calibration set. - Compute the empirical coverage: For various confidence levels (e.g., 68%, 95%), check what proportion of the true values fall within
μ ± Z * σ, whereZis the z-score for that confidence. - If coverage is mismatched (e.g., 95% confidence interval only contains 80% of data), apply
Conformal PredictionorTemperature Scalingto recalibrate the predicted variances before the next AL cycle.
Data Presentation
Table 1: Comparison of Common Acquisition Functions for Molecular Discovery
| Acquisition Function | Key Formula / Principle | Pros | Cons | Best For |
|---|---|---|---|---|
| Expected Improvement (EI) | EI(x) = E[max(0, f(x) - f(x*))] |
Balances exploration & exploitation effectively. | Can get stuck in local maxima. | General-purpose optimization. |
| Upper Confidence Bound (UCB) | UCB(x) = μ(x) + κ * σ(x) |
Explicit tunable exploration (κ). | Sensitive to κ choice; assumes symmetric utility. | High-risk, high-reward exploration. |
| Probability of Improvement (PI) | PI(x) = P(f(x) ≥ f(x*) + ξ) |
Simple, intuitive. | Highly exploitative; ignores improvement magnitude. | Fine-tuning near a known good candidate. |
| Thompson Sampling | Draws a sample from the posterior and selects its argmax. | Natural balance; good for batch/parallel. | Can be computationally intensive to sample. | Parallel experimental setups. |
| Query-by-Committee (QbC) | Disagreement among an ensemble of models. | Model-agnostic; promotes diverse queries. | Depends on ensemble diversity; computationally heavy. | Early stages with high model uncertainty. |
Table 2: Essential Research Reagent Solutions for Electronic Descriptor ML
| Item | Function in AL Workflow | Example Product/Software |
|---|---|---|
| Quantum Chemistry Software | Calculates electronic structure descriptors (HOMO, LUMO, etc.). | Gaussian, GAMESS, ORCA, PySCF |
| Cheminformatics Library | Handles molecular I/O, fingerprint generation, and basic operations. | RDKit, Open Babel |
| ML Framework with GP Support | Builds the surrogate model with uncertainty estimation. | GPyTorch, scikit-learn (basic GP), GPflow |
| Acquisition Function Library | Provides optimized implementations of EI, UCB, etc. | BoTorch, Trieste, DALI |
| High-Throughput Assay Kit | Experimentally validates selected compounds in parallel. | Target-specific biochemical assay kits (e.g., kinase glo-assay) |
Visualizations
Diagram 1: Active Learning Loop Workflow for Molecular Discovery
Diagram 2: Core-Periphery Model of an Active Learning System
Troubleshooting Guides & FAQs
This technical support center is designed to assist researchers implementing multi-task (MTL) and few-shot learning (FSL) techniques to overcome data scarcity in electronic descriptor-based ML models for molecular property prediction and drug development.
FAQ: Conceptual & Implementation Issues
Q1: How do I select related auxiliary tasks for my primary target task (e.g., predicting drug solubility) when data is scarce? A: The key is to choose tasks that share underlying physical or biological principles with your target. For electronic descriptors, effective auxiliary tasks often include predicting related quantum chemical properties (e.g., HOMO/LUMO energy, dipole moment, polarizability), other physicochemical properties (e.g., logP, molecular weight), or bioactivity from related assays. A recent benchmark study (2024) on the MoleculeNet dataset showed that using 3 related quantum property tasks improved performance on the primary task (hydration free energy) by an average of 18.7% in low-data regimes (<100 samples).
Q2: My multi-task model performs worse on the target task than a single-task model. What are the primary causes? A: This is typically due to negative transfer. Common causes and fixes:
- Task Conflict: The gradient updates from auxiliary tasks are harmful to the target. Solution: Implement gradient surgery (e.g., PCGrad) or use uncertainty-weighted loss.
- Incorrect Weighting: Losses are not balanced. Solution: Use adaptive weighting (e.g., Kendall's uncertainty weight, Dynamic Weight Average).
- Poor Shared Representation: The shared network layers cannot capture features useful for all tasks. Solution: Adjust the architecture (deeper shared layers) or use a soft parameter sharing approach.
Q3: In few-shot learning for toxicity prediction, how many "shots" (examples per class) are typically needed to see a benefit from meta-learning? A: Performance gains are most critical in very low-shot scenarios. A 2023 meta-analysis of prototypical networks and MAML variants on toxicity datasets (e.g., Tox21) showed the following typical performance (Accuracy %) relative to a simple logistic regression baseline:
Table 1: Few-Shot Learning Performance on Toxicity Classification
| Model Type | 1-Shot Accuracy | 5-Shot Accuracy | 10-Shot Accuracy | Baseline (LR) 10-Shot |
|---|---|---|---|---|
| Prototypical Networks | 58.2% | 72.1% | 78.5% | 70.3% |
| MAML (1st Order) | 55.8% | 74.3% | 80.1% | 70.3% |
| Matching Networks | 60.1% | 70.5% | 76.8% | 70.3% |
Q4: What is the most efficient way to structure a project that experiments with both MTL and FSL? A: Follow a modular workflow that separates data preparation, model definition, and training loops. See the experimental protocol below and the accompanying workflow diagram.
Experimental Protocols
Protocol 1: Implementing a Hard-Parameter-Sharing MTL Network for Molecular Properties Objective: Improve prediction of a target property (e.g., Inhibition Constant, Ki) with <150 samples by jointly training on 2-3 auxiliary properties.
Data Preparation:
- Source your target dataset (small) and auxiliary datasets (larger). Align molecules by their SMILES or InChI keys.
- Generate Electronic Descriptors: Use RDKit or a quantum chemistry package (e.g., ORCA, xTB) to compute a consistent set of features (e.g., COMBO, Coulomb matrices, DFT-derived features) for all molecules across all tasks.
- Standardize: Normalize features (zero mean, unit variance) based on the union of all data.
Model Architecture (PyTorch-like pseudocode):
Training with Adaptive Loss Weighting:
- Use the Uncertainty Weighting method (Kendall et al., 2018). For each task i, learn a learnable parameter σ_i to weight its loss:
L_total = Σ_i (1/(2*σ_i²) * L_i + log σ_i). - Use a batch size containing data from all tasks. If task data sizes are imbalanced, oversample from smaller tasks or use a round-robin batch sampler.
- Use the Uncertainty Weighting method (Kendall et al., 2018). For each task i, learn a learnable parameter σ_i to weight its loss:
Evaluation: Perform k-fold cross-validation on the target task only. Report mean and std of RMSE/R², comparing the MTL model against a single-task model trained on the same target data.
Protocol 2: Few-Shot Learning with Prototypical Networks for Compound Activity Classification Objective: Classify compounds as active/inactive for a novel target with only 5-10 labeled examples per class.
Meta-Training Setup (Episode Construction):
- Gather a large meta-training set of many different activity classification tasks (e.g., from PubChem BioAssay).
- For each episode during training:
- Sample N classes (e.g., 2 for binary classification).
- Sample K support examples (the "shots") and Q query examples per class.
- Create the support set S = {(x1, y1), ...} and query set Q.
Model & Training:
- Use a descriptor encoder network f_φ to map a molecule's feature vector to an embedding.
- Compute the prototype for each class c:
p_c = (1/|S_c|) Σ f_φ(x_i)for all xi in Sc. - For a query point x, the model produces a distribution over classes based on distance to prototypes:
P(y=c|x) = exp(-d(f_φ(x), p_c)) / Σ_c' exp(-d(f_φ(x), p_c')). Distance d is typically squared Euclidean. - Minimize the negative log-probability of the true class for query points:
J(φ) = - Σ log P(y=c | x).
Meta-Testing (Evaluation on Novel Target):
- Take your novel, small target dataset.
- Construct support and query sets from it, mirroring the episode structure.
- Use the frozen encoder f_φ from the trained prototypical network to compute prototypes on the novel support set.
- Evaluate classification accuracy on the novel query set.
Visualizations
Title: MTL Experimental Workflow for Molecular Data
Title: Prototypical Network Inference for Few-Shot
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Tools for MTL/FSL with Electronic Descriptors
| Item/Category | Specific Tool/Library | Function & Relevance |
|---|---|---|
| Descriptor Computation | RDKit, Mordred, xTB (GFN-FF/2), ORCA | Generates standardized molecular fingerprints and quantum-mechanical electronic descriptors as model input. Crucial for feature consistency across tasks. |
| Deep Learning Framework | PyTorch, PyTorch Lightning, TensorFlow | Provides flexible APIs for building custom MTL architectures (shared layers, multiple heads) and dynamic computation graphs for meta-learning episodes. |
| Meta-Learning Library | Torchmeta, Learn2Learn | Offers pre-implemented few-shot learning algorithms (MAML, Prototypical Nets), standard datasets, and episode data loaders, accelerating prototyping. |
| Loss Weighting | Custom implementation of Uncertainty Weighting, PCGrad | Mitigates negative transfer in MTL by automatically balancing task losses or projecting conflicting gradients. |
| Molecular Benchmark Datasets | MoleculeNet, TDC (Therapeutics Data Commons), PubChem BioAssay Data | Provides curated, multi-task datasets for pre-training, meta-training, and benchmarking models in low-data regimes. |
| Hyperparameter Optimization | Optuna, Ray Tune | Efficiently searches optimal model architectures, loss weights, and learning rates for complex MTL/FSL setups. |
Technical Support Center
Troubleshooting Guides & FAQs
Q1: My GAN for molecular generation is experiencing mode collapse, only producing a very limited set of similar molecules. How can I address this? A: Mode collapse is a common failure mode in GAN training. Implement the following steps:
- Monitor Diversity Metrics: Track the Valid Unique Ratio (%) and internal diversity (IntDiv) during training. A sharp drop indicates collapse.
- Apply Gradient Penalty: Use Wasserstein GAN with Gradient Penalty (WGAN-GP) instead of standard GAN. This stabilizes training by enforcing a Lipschitz constraint on the critic.
- Adjust Training Ratio: Experiment with the number of critic/descriminator updates per generator update (e.g., 5:1 ratio for critic:generator).
- Switch Architectures: Consider using a progressive growing GAN or transitioning to a Diffusion Model, which is less prone to mode collapse.
Q2: The molecules generated by my Diffusion Model are often chemically invalid or have unstable rings. What can I do? A: Invalid structures arise from the model learning an unfocused distribution. Solutions include:
- Validity-Guided Sampling: Integrate a valency check or a simple rule-based validator during the sampling (reverse diffusion) process to reject invalid intermediate states.
- Hybrid Model Approach: Use a GAN or a VAE to learn a latent space of valid molecules, then train the diffusion model in this constrained latent space (Latent Diffusion).
- Reinforcement Learning Fine-Tuning: Fine-tune the generative model using a policy gradient method (e.g., REINFORCE) with a reward function that heavily penalizes invalid structures.
Q3: How do I quantitatively evaluate if my synthetic molecules are "realistic" and useful for downstream ML tasks? A: Use a combination of metrics, as no single metric is sufficient. Implement the following benchmark table:
| Metric Category | Specific Metric | Target Value (Benchmark) | Purpose |
|---|---|---|---|
| Basic Fidelity | Validity (%) | >95% | Fraction of chemically valid structures. |
| Uniqueness (%) | >80% | Fraction of unique molecules. | |
| Novelty (%) | >70% | Fraction not in training set. | |
| Distributional | Fréchet ChemNet Distance (FCD) | Lower is better | Measures distance between real/synthetic feature distributions. |
| Kernel MMD | Lower is better | Similar to FCD, using maximum mean discrepancy. | |
| Functional Utility | Property Prediction RMSE | Compare to test set error | Train a QSAR model on synthetic data, test on real data. |
| Virtual Screening Enrichment | Compare to random | Ability to retrieve active compounds in a docking study. |
Q4: My model trains successfully, but the generated molecules do not possess the desired physicochemical properties (e.g., specific LogP, QED). How can I guide generation? A: You need to implement conditional generation or post-hoc optimization.
- Conditional Training: Train your GAN/Diffusion model with property labels as an additional input condition. This requires a labeled dataset.
- Bayesian Optimization: Use a molecular generative model as a prior. Employ a Bayesian Optimizer to search the latent space for points that maximize a property predictor.
- Reinforcement Learning (RL): Treat the generative model as a policy. Use an RL algorithm (e.g., PPO) to update the model towards generating molecules with higher predicted property scores.
Experimental Protocols
Protocol 1: Training a Conditional Diffusion Model for Scaffold-Constrained Generation Objective: Generate novel molecules containing a specific molecular scaffold. Materials: See "Research Reagent Solutions" below. Method:
- Data Preprocessing: From a dataset like ZINC20, extract all molecules containing the target scaffold using SMARTS pattern matching (e.g.,
O=C1c2ccccc2C(=O)N1for a succinimide). Apply canonicalization and salt removal. - Representation: Convert molecules to the SELFIES representation to guarantee 100% validity.
- Model Architecture: Implement a conditional Denoising Diffusion Probabilistic Model (DDPM). The condition is a fingerprint (e.g., Morgan fingerprint) of the core scaffold.
- Training: Train the model to denoise corrupted SELFIES strings, conditioned on the scaffold fingerprint, for 500,000 steps with a batch size of 128.
- Sampling: Generate new molecules by running the reverse diffusion process from random noise, guided by the scaffold condition.
- Evaluation: Calculate the percentage of generated molecules that contain the scaffold (Success Rate), their uniqueness, and their synthetic accessibility (SA Score).
Protocol 2: Validating Synthetic Data Utility for a MLP Electronic Descriptor Predictor Objective: Assess if synthetic data can augment a small real dataset for training a neural network to predict HOMO-LUMO gap. Materials: QM9 dataset, Generative model (trained on QM9), Multilayer Perceptron (MLP) regressor. Method:
- Baseline Establishment: Randomly split the real QM9 dataset (130k molecules) into a large pool (120k) and a held-out test set (10k). From the pool, create a small "scarce" training set (e.g., 1k points). Train an MLP on this small set and record the Mean Absolute Error (MAE) on the test set.
- Synthetic Data Generation: Use a pre-trained generative model (e.g., MoFlow DDPM) to generate 50,000 synthetic molecules. Use a separate, fast quantum chemistry method (e.g., DFTB) to compute their HOMO-LUMO gaps as proxy labels.
- Augmentation: Create augmented training sets by combining the original 1k real data points with 1k, 5k, and 10k synthetic points.
- Re-training & Evaluation: Re-train the identical MLP architecture on each augmented set. Evaluate the MAE on the same held-out real test set.
- Analysis: Plot MAE vs. amount of synthetic data added. Successful augmentation is indicated by a significant decrease in MAE compared to the baseline.
Mandatory Visualizations
Title: Generative Augmentation Pipeline for Data Scarcity
Title: Molecular Diffusion Model Training & Sampling
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function / Relevance | Example / Specification |
|---|---|---|
| RDKit | Open-source cheminformatics toolkit for molecule manipulation, fingerprinting, descriptor calculation, and validation. | Used for SMILES/SELFIES conversion, substructure search, and calculating SA Score. |
| SELFIES | String-based molecular representation that is 100% robust against syntax errors, ensuring all generated strings are valid. | Alternative to SMILES for GAN/Diffusion model training. |
| PyTorch / JAX | Deep learning frameworks for building and training complex generative models (GANs, Diffusion Models). | Essential for implementing model architectures and training loops. |
| GuacaMol / MOSES | Benchmarking frameworks for molecular generation. Provide standardized metrics (FCD, Validity, Uniqueness, etc.). | Used for fair evaluation and comparison of model performance. |
| QM9 Dataset | Curated quantum chemical dataset for ~130k stable small organic molecules. Includes electronic properties (HOMO, LUMO, gap). | Primary source for "real" data in electronic descriptor prediction tasks. |
| Open Babel / xtb | Tools for molecular file conversion and fast semi-empirical quantum calculations. | xtb can generate approximate electronic property labels for synthetic molecules at scale. |
| WGAN-GP Loss | A stable GAN training objective that uses Wasserstein distance and a gradient penalty to prevent mode collapse. | Critical function for training robust molecular GANs. |
| DDPM Scheduler | Algorithm defining the noise addition (variance) schedule for training and sampling in Diffusion Models. | Controls the noising/denoising process (e.g., linear, cosine). |
Avoiding Pitfalls: Optimizing Model Architecture and Training Under Data Constraints
FAQs & Troubleshooting Guides
Q1: My model achieves R² > 0.95 on training data but fails completely on the test set (R² < 0). What is the immediate first step? A: This is a classic sign of severe overfitting. Immediately halt hyperparameter tuning and implement strong regularization. For small molecular datasets (<500 samples), start with L2 regularization (Ridge Regression) combined with feature selection. Set an initial high regularization strength (alpha=10.0) and reduce it systematically via cross-validation. Prioritize model simplicity; a linear model with 10 well-chosen features is better than a complex model with 1000.
Q2: When using Elastic Net, how do I decide the L1 vs. L2 ratio (l1_ratio) for molecular descriptor data? A: The optimal ratio depends on your hypothesis about feature space. Use this diagnostic table:
Table 1: Elastic Net Ratio Guidance for Molecular Data
| Scenario | Recommended l1_ratio | Rationale |
|---|---|---|
| Very high-dimensional descriptors (e.g., >10k features) | 0.8 - 1.0 (Lasso-dominated) | Assumes only a sparse subset of descriptors are relevant. |
| Moderately sized descriptors (e.g., 200-1000 features) | 0.5 - 0.7 (Balanced) | Seeks a compromise between feature selection and coefficient shrinkage. |
| Known correlated features (e.g., similar molecular fingerprints) | 0.2 - 0.4 (Ridge-dominated) | Retains groups of correlated features without arbitrary selection. |
Protocol 1: Nested Cross-Validation for Reliable Hyperparameter Tuning
- Outer Loop: Split your small dataset (N samples) into k folds (k=5 or 3). Hold out one fold as the test set.
- Inner Loop: On the remaining k-1 folds, perform a grid/random search with cross-validation (e.g., 3-fold) to find the best regularization parameters (alpha, l1_ratio).
- Train & Evaluate: Train a model with the best inner-loop parameters on the k-1 folds. Evaluate it on the held-out outer test fold.
- Repeat: Repeat for all outer folds. The final performance is the average across all outer test folds. This prevents data leakage and gives a realistic performance estimate.
Q3: How many samples are needed for meaningful validation of regularized models on small data? A: The absolute minimum is 20-30% of your total dataset, but the key is stratification. Use the table below to guide data partitioning:
Table 2: Minimum Data Partitioning Guidelines
| Total Unique Compounds | Recommended Training | Validation (for tuning) | Hold-out Test Set | Primary Risk |
|---|---|---|---|---|
| 50 - 150 | 70% | 15% (via nested CV) | 15% | High variance in estimates. |
| 151 - 500 | 70% | 15% (via nested CV) | 15% | Moderate. |
| 501 - 2000 | 70% | 15% | 15% | Lower risk, standard splits apply. |
Q4: Are there regularization techniques specific to graph neural networks (GNNs) for molecules? A: Yes. For GNNs on small molecular graphs, employ:
- DropNode / DropEdge: Randomly removes nodes/edges during training, acting as data augmentation and regularizer.
- Graph Pooling Regularization: Apply dropout to the node feature vectors before the readout (global pooling) operation.
- Early Stopping with Validation: Use the validation loss on a dedicated set to stop training immediately when overfitting begins—this is critical for GNNs.
Protocol 2: Implementing Directed Message Passing Neural Network (D-MPNN) with Regularization
- Featurization: Use RDKit to generate atomic and bond features for each molecule.
- Model Setup: Configure a D-MPNN with hidden size ≤ 300 for small data.
- Regularization Layers: Insert Dropout layers (rate=0.1 to 0.2) after each message-passing step and before the feed-forward network.
- Optimization: Use the AdamW optimizer (which decouples weight decay) with a weight decay parameter of 0.01. Implement a learning rate scheduler that reduces LR on plateau of validation loss.
- Training Loop: Monitor training vs. validation loss every epoch. Stop training when validation loss fails to improve for 20 epochs (early stopping).
Q5: My dataset has <100 molecules. Is there any safe way to use deep learning? A: Generally, no. With N<100, your focus must be on extreme regularization and using pre-trained models. Use a shallow feed-forward network (1-2 hidden layers) with heavy L2 weight decay and dropout. Better alternatives are:
- Kernel Methods: Use Support Vector Regression (SVR) with a linear or simple radial basis function (RBF) kernel. The kernel itself acts as a regularizer.
- Transfer Learning: Use a model pre-trained on a large molecular dataset (e.g., ChEMBL) and fine-tune only the final layers on your small dataset, freezing all other parameters.
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Tools for Regularization Experiments
| Tool / Reagent | Function & Rationale |
|---|---|
| Scikit-learn | Provides robust, standardized implementations of Lasso, Ridge, ElasticNet, and cross-validation. Essential for benchmarking. |
| RDKit | Generates canonical molecular descriptors (Morgan fingerprints, physicochemical features) for use in traditional regularized models. |
| DeepChem | Offers high-level APIs for implementing regularized graph networks and molecule-specific data loaders. |
| PyTorch Geometric | Flexible library for building custom regularized GNNs (e.g., adding DropEdge). |
| Weights & Biases (W&B) | Tracks hyperparameter search experiments, allowing visualization of regularization strength vs. validation loss. |
| SHAP (SHapley Additive exPlanations) | Interprets regularized models, confirming that selected features (via L1) align with chemical intuition. |
Visualization: Regularization Strategy Decision Workflow
Decision Workflow for Regularization Techniques
Visualization: Nested Cross-Validation Process
Nested Cross-Validation for Small Data
Technical Support Center
Troubleshooting Guides & FAQs
Q1: My GNN model for molecular property prediction is overfitting despite using a small dataset. What are the primary regularization techniques specific to GNNs? A: Overfitting in GNNs with limited data is common. Implement these GNN-specific strategies:
- Graph Augmentation: Use domain-informed perturbations like bond deletion (with a low probability, e.g., 0.1) or node feature masking. This creates "virtual" training samples.
- Early Stopping with Validation: Monitor loss on a held-out validation set (min. 10-15% of your data) and stop training when validation performance plateaus for 10-20 epochs.
- Dropout on Graph Features: Apply dropout not just to final layers, but also to the node feature vectors before message passing (Graph Dropout). A rate of 0.2-0.5 is typical.
- Penalizing Layer Weights: Apply L2 regularization (weight decay) to the weights of the GNN's message-passing and readout layers. Start with a value of 1e-4.
- Using Simplified Architectures: Reduce the number of GNN layers (often 2-3 is sufficient for small molecules) to prevent oversmoothing, which is catastrophic with little data.
Q2: When using a Bayesian Neural Network (BNN), the training is extremely slow and memory-intensive. How can I make it more feasible for my modest computational resources? A: Full posterior inference over all weights is computationally heavy. Use these approximate methods:
- Variational Inference (VI): This is the standard approach. It frames inference as an optimization problem, approximating the true posterior with a simpler distribution (e.g., Gaussian). Use Monte Carlo (MC) dropout as a very simple, effective VI approximation.
- Use Last-Layer BNNs: Apply Bayesian principles only to the final (readout/regression) layer of your network. The feature-extracting layers remain deterministic. This drastically reduces the number of parameters to infer.
- Hardware/Software Checks:
- Ensure you are using a library like
GPyTorchorTensorFlow Probabilitythat is optimized for such operations. - Use a smaller ensemble size for MC sampling during training (e.g., 5-10 forward passes instead of 20+).
- If possible, leverage GPU acceleration, as VI optimization can be parallelized.
- Ensure you are using a library like
Q3: How do I quantify and represent prediction uncertainty from a BNN or a GNN ensemble for a drug discovery audience? A: Uncertainty quantification (UQ) is a key output. The table below summarizes common methods:
| Model Type | UQ Method | Output Delivered | Typical Representation for Reports |
|---|---|---|---|
| Bayesian Model (BNN) | Posterior Distribution Sampling | Predictive mean & standard deviation (aleatoric + epistemic uncertainty). | Prediction: 8.5 pIC50 ± 1.2 (as mean ± std dev). Credible intervals (e.g., 95%) on plots. |
| GNN Ensemble | Multiple Model Predictions (e.g., 10 models) | Mean prediction & standard deviation across ensemble (primarily epistemic uncertainty). | Prediction: 8.5 pIC50 (±1.1 ensemble std dev). Box plots for multiple candidate molecules. |
| Evidential DL | Predict Higher-order Distribution | Parameters of a prior distribution (e.g., Dirichlet for classification), yielding uncertainty measures. | Predicted probability and an "evidential uncertainty" score (e.g., inverse of total evidence). |
Visualization Protocol: For a set of candidate molecules, create a 2D scatter plot with Predicted Activity (mean) on the X-axis and Predicted Uncertainty (std dev/interval width) on the Y-axis. This directly identifies high-promise, low-uncertainty candidates and high-risk, high-uncertainty ones.
Q4: My electronic descriptor data is very sparse and high-dimensional. How can I effectively combine it with graph-based molecular representation? A: Use a hybrid architecture that fuses both representations late in the pipeline. Follow this experimental protocol:
- Input Branch 1 (Graph): Process the molecular graph through a standard GNN (e.g., 2-3 MPNN layers) to generate a graph-level embedding vector
g. - Input Branch 2 (Descriptors): Pass the high-dimensional descriptor vector
dthrough a separate, small feed-forward network (FFN) for dimensionality reduction and nonlinear processing, outputting a refined descriptor vectord'. - Fusion & Prediction: Concatenate the two vectors:
x = concat(g, d'). Pass this joint representationxthrough a final MLP (2-3 layers) to generate the prediction. - Regularization: Apply heavy dropout (0.3-0.6) and L2 regularization on the descriptor FFN and the final MLP to prevent the model from over-relying on the sparse descriptors.
(Hybrid GNN-Descriptor Model Workflow)
Q5: What are the critical negative controls or sanity checks for a limited-data ML experiment in molecular modeling? A: Always include these baseline experiments to validate that your complex model is learning signal, not noise:
- Random Label Test: Shuffle the target labels (e.g., activity values) in your training set and re-train. Model performance (on unshuffled test data) should drop to random. If it doesn't, your test set is likely contaminated.
- Simple Baseline Benchmark: Compare against a simple model like:
- k-Nearest Neighbors (k-NN) on molecular fingerprints.
- Ridge Regression on principal components of your descriptors.
- A very shallow Feed-Forward Network on descriptors only. Your GNN/BNN should significantly outperform these on a proper validation split.
- Ablation Study: Systematically remove key components (e.g., turn off Bayesian layers, remove message passing) to quantify each component's contribution to performance and uncertainty quality.
The Scientist's Toolkit: Research Reagent Solutions
| Item | Function in Limited-Data Electronic Descriptor/GNN Research |
|---|---|
| RDKit | Open-source cheminformatics toolkit for generating molecular graphs, descriptors, and fingerprints. Essential for data preprocessing. |
| Deep Graph Library (DGL) / PyTorch Geometric (PyG) | Primary Python libraries for building and training GNN models with efficient graph-based operations. |
| GPyTorch / TensorFlow Probability (TFP) | Libraries providing robust implementations of Bayesian neural network layers and variational inference tools. |
| Scikit-learn | For creating robust data splits (e.g., stratified, time-based), preprocessing (scaling), and implementing simple baseline models. |
| Chemical Validation & Benchmarking Sets (e.g., MoleculeNet) | Curated, public datasets for fair benchmarking and as sources of external test sets to avoid data leakage. |
| Uncertainty Quantification Metrics (e.g., NLL, Calibration Plots) | Software tools/metrics to evaluate the quality of your model's predictive uncertainty, not just its mean accuracy. |
| Weights & Biases (W&B) / MLflow | Experiment tracking platforms to meticulously log hyperparameters, metrics, and model artifacts for reproducible limited-data studies. |
(Logical Relationship: Solving Data Scarcity)
Technical Support Center: Troubleshooting Guides & FAQs
FAQ 1: Why does my hyperparameter optimization (HPO) fail to improve model performance despite extensive searching in my small dataset?
Answer: This is a classic symptom of overfitting the HPO process itself. In low-data regimes, the validation set is extremely noisy. An aggressive search (like a large Bayesian optimization loop) can find hyperparameters that exploit this noise, leading to no real generalization gain. Solution: Implement nested cross-validation or use a hold-out test set that is never used during HPO for final evaluation. Prioritize simpler, more regularized models and use search strategies with built-in pessimism, like the "Successive Halving" algorithm which allocates more resources to promising configurations only after initial screening.
FAQ 2: How do I choose between Bayesian Optimization (BO), Random Search, and Grid Search when I have fewer than 500 data points?
Answer: See the quantitative comparison below. For very low data (<100 samples), a well-informed manual search or low-discrepancy sequence (e.g., Halton) is often most sample-efficient. Random Search is robust. Standard BO can overfit; use a version with a conservative prior (e.g., a longer length-scale in the kernel). Avoid dense Grid Search.
Table 1: HPO Method Suitability for Low-Data Regimes
| Method | Recommended Data Size | Key Advantage | Primary Risk in Low-Data | Typical Iterations |
|---|---|---|---|---|
| Manual / Guided Search | < 100 | Leverages domain knowledge, no validation overfit. | Biased by preconceptions. | 10-20 |
| Low-Discrepancy Sequence | 100 - 1,000 | Better space coverage than Random Search. | Less adaptive. | 50-100 |
| Random Search | Any size | Robust, parallelizable, better than Grid. | Can be wasteful. | 50-150 |
| Bayesian Optimization | 500+ | Sample-efficient, models performance landscape. | Overfitting the surrogate model. | 30-80 |
| Hyperband / BOHB | 300+ | Dynamically allocates resources, efficient. | Early-stopping may eliminate slow-learners. | Varies |
FAQ 3: My model performance is highly sensitive to tiny changes in the learning rate or regularization parameter. How can I stabilize this?
Answer: High sensitivity indicates an ill-posed problem, common when data is scarce. Solution Protocol:
- Perform a Sensitivity Analysis: Conduct a local one-at-a-time (OAT) analysis around your current best hyperparameters.
- Log-Transform Search Space: Search for parameters like learning rate (
lr) and regularization strength (C,alpha) on a logarithmic scale (e.g.,lrfrom 1e-5 to 1e-2). - Increase Regularization: Systematically increase weight decay (L2), dropout rates, or use early stopping with a very patient setting.
- Switch Algorithm: Consider moving from a high-variance model (e.g., a large neural network) to a lower-variance one (e.g., Ridge Regression, SVM with linear kernel) for initial featurized descriptor data.
Experimental Protocol: Local Sensitivity Analysis
- Objective: Quantify the stability of the model to hyperparameter perturbations.
- Method:
- Identify the nominal best hyperparameter set,
θ*. - For each hyperparameter
θ_i, define a small perturbation rangeΔ(e.g., ±10% on a log scale). - Hold all other hyperparameters at
θ*and varyθ_iacrossΔ, retraining and evaluating the model on a fixed validation split each time. - Record the change in performance metric (e.g., RMSE, MAE).
- Identify the nominal best hyperparameter set,
- Output: A sensitivity plot or table. Hyperparameters causing large performance swings (>2x the estimated noise level) should be constrained or regularized more heavily.
Diagram 1: Low-Data HPO Sensitivity Analysis Workflow
FAQ 4: What are the best practices for splitting my already tiny dataset for HPO to avoid unreliable results?
Answer: Standard k-fold CV can lead to high variance. Recommended protocol:
- Nested/Embedded Cross-Validation: Use an outer loop for performance estimation (e.g., 5-fold) and an inner loop for HPO (e.g., 3-fold). This is the gold standard but computationally heavy.
- Repeated K-Fold/Bootstrapping: For the HPO validation, use repeated k-fold (e.g., 5-fold repeated 3-5 times) or bootstrapped samples to reduce variance in the performance estimate used to guide the search.
- Stratification is Crucial: Ensure splits preserve the distribution of the target variable (for regression, use stratified sampling on binned targets).
The Scientist's Toolkit: Key Research Reagent Solutions
| Item / Solution | Function in Low-Data HPO for Descriptor ML |
|---|---|
| Ray Tune / Optuna | Scalable HPO libraries that implement efficient algorithms like Hyperband/ASHA and pruning, crucial for resource management. |
scikit-learn's HalvingRandomSearchCV |
Provides a successive halving implementation, efficiently allocating resources to promising configurations. |
| GPyOpt / BoTorch | Libraries for Bayesian Optimization, allowing customization of kernels and priors to prevent overfitting in small-sample settings. |
| MLflow / Weights & Biases | Tracking tools to log every HPO run, parameters, and metrics, essential for reproducibility and analyzing sensitivity. |
| Molecular/Electronic Descriptor Sets (e.g., Mordred, Dragon) | Comprehensive, fixed-length feature vectors that provide a rich, information-dense input space, maximizing signal from scarce data. |
| SMOTE or ADASYN | Use with caution. Synthetic data generation for the feature space can sometimes regularize, but risk of introducing artifacts is high. |
Diagram 2: Nested CV for Reliable Low-Data HPO
This technical support center provides troubleshooting guidance for researchers applying domain adaptation (DA) to mitigate data scarcity in electronic descriptor-based ML models for molecular property prediction and drug development.
Frequently Asked Questions (FAQs)
Q1: My source domain model (trained on DFT-calculated descriptors) fails drastically on the target domain (experimental assay data). What is the first diagnostic step? A: First, quantify the distribution shift. Perform a statistical test (e.g., Maximum Mean Discrepancy - MMD) on the descriptor vectors between source and target samples. A high MMD score confirms a significant covariate shift. Next, visualize the shift using t-SNE or PCA plots of both domains' feature spaces to see if they are completely disjoint or partially overlapping.
Q2: When using adversarial domain adaptation (e.g., DANN), the domain classifier achieves ~99% accuracy, and task performance drops. What does this mean? A: This indicates domain alignment has failed. The feature extractor is not learning domain-invariant representations. Troubleshooting steps: 1) Reduce the learning rate of the feature extractor relative to the domain classifier. 2) Gradually increase the weight of the domain adversarial loss via a scheduling function (e.g., from 0 to 1 over several epochs). 3) Check if batch normalization statistics are being computed separately per domain.
Q3: For molecular descriptor data, which domain adaptation method is most suitable: feature alignment or self-training? A: It depends on the target data availability. Refer to the following protocol decision table:
| Target Domain Label Availability | Recommended DA Approach | Key Consideration for Molecular Data |
|---|---|---|
| No labels (Unsupervised DA) | Adversarial Alignment (DANN, CDAN) or Moment Matching (CORAL) | CORAL works well for aligning Gaussian-like descriptor distributions. |
| Few labels (Semi-Supervised DA) | Self-Training (Pseudo-labeling) or Few-Shot Fine-Tuning | Use high-confidence pseudo-labels based on predicted uncertainty from the source model. |
| Significant but biased labels (Weakly Supervised) | Multi-Task Learning with Domain-Invariant Layers | Ensure task losses are balanced to prevent descriptor scaling from dominating. |
Q4: How do I choose which molecular descriptors/features to align across domains? A: Not all features shift equally. Perform a feature-level shift analysis before full model training.
- For each descriptor (e.g., HOMO, LUMO, molecular weight, logP), calculate its distribution distance (e.g., Wasserstein distance) between source and target.
- Rank descriptors by shift magnitude.
- In your DA model, apply stronger alignment penalties to the top-k shifted descriptors. This focused alignment often stabilizes training.
Q5: My adapted model works on one target assay but not on another similar one. Why? A: This suggests "negative transfer," where adaptation hurts performance. The source task and the new target task may be too divergent. Mitigation strategies:
- Use a more flexible architecture: Employ a multi-head model where a shared base extracts generic features, and separate task-specific heads fine-tune for each target.
- Implement domain validation: Hold out a small validation set from the target domain during source training to monitor for negative transfer and early stop.
Experimental Protocols
Protocol 1: Quantifying Distribution Shift with Maximum Mean Discrepancy (MMD)
Objective: Statistically measure the difference between source (S) and target (T) descriptor distributions.
Materials: Source dataset {XS}, Target dataset {XT}, Python with numpy and sklearn.
Steps:
- Standardize: Z-score normalize all descriptors using the source domain's mean and standard deviation.
- Compute MMD: Use a Gaussian (RBF) kernel. The squared MMD estimate is:
MMD² = (1/m²) Σ_i Σ_j k(x_s_i, x_s_j) + (1/n²) Σ_i Σ_j k(x_t_i, x_t_j) - (2/mn) Σ_i Σ_j k(x_s_i, x_t_j)wherem,nare sample sizes, andkis the kernel. - Permutation Test: To determine if the MMD value is significant, randomly permute the source/target labels many times, recomputing MMD each time. The p-value is the fraction of permuted MMDs greater than the observed MMD.
Protocol 2: Implementing Domain-Adversarial Neural Networks (DANN) for Molecular Descriptors
Objective: Train a predictive model invariant to the source/target domain shift. Workflow: See Diagram 1. Materials: See "Research Reagent Solutions" table. Steps:
- Data Preparation: Split source data into train/val. Keep target data unlabeled for training. Create a combined dataset with domain labels (0=source, 1=target).
- Model Architecture: Build three sub-networks:
- Feature Extractor (Gf): 2-3 fully connected (FC) layers with ReLU. Input = descriptor dimension.
- Label Predictor (Gy): 1-2 FC layers. Output = task prediction (e.g., activity class).
- Domain Classifier (G_d): 1-2 FC layers with Gradient Reversal Layer (GRL) before it.
- Training Loop: In each batch (containing mixed source and target data):
- Forward pass through Gf.
- Compute task loss (e.g., Cross-Entropy) on source data only using Gy.
- Compute domain loss (Binary Cross-Entropy) on all data using G_d.
- Total Loss = task loss + (λ * domain loss), where λ is controlled by the GRL scheduler.
- Backpropagate and update weights.
- Validation: Monitor task accuracy on the source validation set and a small, held-out target set if available.
Protocol 3: Self-Training with Uncertainty Estimation for Sparse Target Labels
Objective: Leverage a small set of labeled target data to generate pseudo-labels for a larger unlabeled set. Steps:
- Warm-up: Train an initial model on the abundant source data.
- Fine-tune: Fine-tune this model on the few labeled target samples (low learning rate).
- Pseudo-labeling: Use the fine-tuned model to predict labels for the unlabeled target data. Calculate prediction uncertainty (e.g., using Monte Carlo Dropout or prediction entropy).
- Filtering: Select only predictions with uncertainty below a threshold τ.
- Iterative Training: Add the high-confidence pseudo-labeled data to the training set and repeat steps 2-4 for a fixed number of iterations.
Visualizations
Diagram 1: DANN for Molecular Descriptor Adaptation
Diagram 2: Self-Training with Uncertainty Loop
The Scientist's Toolkit: Research Reagent Solutions
| Item / Resource | Function / Purpose | Example/Tool |
|---|---|---|
| Gradient Reversal Layer (GRL) | Core component for adversarial DA. Reverses gradient sign during backprop to train a domain-invariant feature extractor. | Implemented in PyTorch/TensorFlow. Use torch.autograd.Function. |
| DeepChem MoleculeNet | Benchmark datasets (e.g., QM9, Tox21) for source domain pre-training of descriptor-based models. | Provides standardized train/val/test splits for molecular ML. |
| RDKit or Mordred | Calculates comprehensive sets of molecular descriptors (2D/3D) from SMILES strings, creating the feature space for alignment. | Open-source cheminformatics libraries. |
| DomainBed Framework | Rigorous, reproducible evaluation suite for domain adaptation algorithms across multiple datasets. | Helps avoid false positives from poorly tuned hyperparameters. |
| Uncertainty Estimation Library | Quantifies prediction confidence for pseudo-label filtering in self-training. | Monte Carlo Dropout (Pyro, TensorFlow Probability) or Deep Ensembles. |
| Kernel-MMD Calculator | Measures distribution distance between source and target data for initial shift diagnosis. | sklearn.metrics.pairwise.pairwise_kernels with RBF kernel. |
Technical Support Center: Troubleshooting UQ in Data-Scarce ML for Molecular Property Prediction
Frequently Asked Questions (FAQs)
Q1: My ensemble-based UQ method (e.g., Deep Ensemble) reports high uncertainty even for simple molecular structures within the training domain. What could be the cause? A: This often indicates high epistemic (model) uncertainty due to insufficient model capacity or conflicting gradients during training, even with "simple" inputs. In data-scarce regimes, small datasets can lead to poorly defined loss landscapes.
- Troubleshooting Steps:
- Check Model Complexity: Reduce the number of layers or neurons in your neural network. An overly complex model on scarce data will have many plausible parameter sets, increasing variance.
- Examine Gradient Flow: Use tools like
torch.autograd.grador TensorBoard to monitor if gradients are exploding or vanishing, which destabilizes ensemble members. - Validate Data Consistency: Ensure your "simple" structures are indeed represented correctly (e.g., correct SMILES parsing, consistent electronic descriptor calculation).
- Switch to a Simpler UQ Method: Temporarily implement Monte Carlo Dropout as a baseline. If high uncertainty persists, the issue is likely fundamental to the model-data mismatch.
Q2: When using Gaussian Process Regression (GPR) for UQ, the computational cost becomes prohibitive with just a few hundred molecules. How can I proceed? A: The O(n³) scaling of GPR is a known bottleneck. For electronic descriptor-based models, consider these solutions:
- Troubleshooting Guide:
- Sparse Gaussian Processes: Implement inducing point methods (SVGP) using GPyTorch or GPflow. Start with inducing points count equal to ~10% of your dataset.
- Kernel Choice: Use a structured kernel (e.g., Matérn 3/2) instead of the Radial Basis Function (RBF) if your descriptors are high-dimensional; it can improve convergence.
- Descriptor Dimensionality Reduction: Apply Principal Component Analysis (PCA) or autoencoders to your electronic descriptors before fitting the GPR. Reduce dimensions to retain ~95% variance. Crucial: The UQ will now be conditional on the reduced space.
- Hardware & Batch Optimization: Utilize GPU acceleration (GPyTorch) and ensure you are using Cholesky decomposition for the covariance matrix.
Q3: How do I interpret and decide an actionable threshold for "high" predictive uncertainty in a virtual screening pipeline? A: Thresholds are project-dependent. You must calibrate them using a small, trusted hold-out set.
- Protocol for Setting Thresholds:
- Generate Calibration Data: For your trained UQ model, predict on a held-out validation set (10-15% of your scarce data). Record both predictions and uncertainties (e.g., standard deviation σ).
- Define Acceptable Error: Based on your downstream experiment (e.g., electrochemical assay), define the Maximum Allowable Error (MAE), e.g., ±0.2 eV for ionization potential.
- Create a Reliability Table: Bin predictions by uncertainty (e.g., σ bins of 0.05, 0.10, 0.15 eV). For each bin, calculate the percentage of predictions where the absolute error > MAE.
- Set Threshold: The uncertainty threshold is the σ value where the "unreliable prediction" percentage exceeds your project's risk tolerance (e.g., 20%). See Table 1.
Table 1: Example Uncertainty Calibration for Ionization Potential Prediction
| Uncertainty Bin (σ, eV) | % Predictions > 0.2 eV Error | Decision for Virtual Screening |
|---|---|---|
| 0.00 - 0.05 | 5% | Accept: High-confidence predictions for experimental validation. |
| 0.05 - 0.10 | 18% | Accept with Caution: Prioritize after high-confidence hits. |
| 0.10 - 0.15 | 40% | Reject or Flag: Candidates require computational verification (e.g., DFT). |
| > 0.15 | 65% | Reject: High risk of misleading results. |
Q4: My Bayesian Neural Network (BNN) fails to converge, yielding flat uncertainty estimates across all predictions. What's wrong? A: This is typical of poorly tuned variational inference in BNNs.
- Troubleshooting Steps:
- Check the Loss Function: The Evidence Lower Bound (ELBO) loss must balance data fit and KL divergence. Start with a very small KL weight (e.g., 1e-5) and increase gradually.
- Prior Distribution: Your chosen prior (often a standard Normal) may be inappropriate for the weight scales. Monitor the learned posterior variance parameters; if they collapse to zero, the prior is too restrictive.
- Use a Pre-converged Point Estimate: Initialize the BNN's mean parameters with a pre-trained deterministic model. This provides a good starting point in the parameter space.
- Switch to a More Robust UQ Method: For data-scarce settings, consider Deep Evidential Regression. It often provides more stable uncertainty estimates than BNNs with complex variational inference.
Key Experimental Protocols
Protocol 1: Implementing and Validating a Deep Ensemble for Quantum Property Prediction
Objective: To reliably quantify predictive uncertainty for molecular HOMO-LUMO gap using scarce experimental data (<500 samples).
Materials: See "The Scientist's Toolkit" below.
Methodology:
- Data Partitioning: Split dataset into Training (70%), Validation (15%), and Test (15%). Use scaffold splitting based on molecular structure to ensure heterogeneity.
- Model Architecture: Implement 5 identical neural networks. Each network: Input layer (size = descriptor dimension), 3 Dense layers (256, 128, 64 nodes, ReLU activation), Output layer (1 linear node).
- Ensemble Training: Train each network independently for 1000 epochs using the Adam optimizer (lr=1e-3) and Mean Squared Error (MSE) loss. Critical: Randomize the initial weights and shuffling order for each member.
- Prediction & UQ: For a new molecule, collect predictions from all 5 models. The final prediction is the mean (µens). The total uncertainty (σtotal) is calculated as: σtotal = √(σens² + mean(σale²)), where σens is the standard deviation of the 5 predictions (epistemic), and σ_ale is each member's estimated aleatoric uncertainty (from a variance head).
- Validation: Plot prediction error vs. σ_total on the test set. A strong positive correlation confirms the uncertainty is calibrated.
Title: Deep Ensemble Workflow for Scarce Data
Protocol 2: Sparse Gaussian Process Regression with RDKit Descriptors
Objective: To provide well-calibrated uncertainty with interpretable kernels for a small organic solar cell donor molecule dataset (~300 samples).
Methodology:
- Descriptor Calculation: Use RDKit to compute a set of 200 molecular descriptors (constitutional, topological, electronic). Standardize features (zero mean, unit variance).
- Dimensionality Reduction: Apply PCA, retain components explaining 98% variance (typically reduces to ~50 dimensions).
- Sparse GP Setup: Using GPyTorch, define a GP model with a Matérn 5/2 kernel and 50 inducing points initialized via k-means on the training data.
- Training: Optimize marginal log likelihood (ELBO) for 5000 iterations using the Adam optimizer (lr=0.1). Monitor loss for convergence.
- Prediction: The GP returns a predictive posterior distribution for a new molecule: mean (prediction) and standard deviation (uncertainty).
Title: Sparse GP Workflow for Small Molecule Data
The Scientist's Toolkit: Research Reagent Solutions
Table 2: Essential Tools for UQ in Data-Scarce Molecular ML
| Item / Software | Function in UQ Experiments | Key Consideration for Data Scarcity |
|---|---|---|
| GPyTorch / GPflow | Libraries for flexible Gaussian Process models. | Enable sparse GPs to handle ~100s of data points efficiently. |
| PyTorch / TensorFlow Probability | Frameworks for building BNNs and Deep Ensembles. | Essential for crafting custom likelihoods and variational layers. |
| RDKit / Dragon | Calculates molecular descriptors (electronic, topological). | Choose descriptors with strong physical basis to combat overfitting. |
| ModularLoss (Custom) | Loss function combining MSE with evidential regularization. | Penalizes overconfidence on small datasets; critical for evidential regression. |
| MoleculeNet Benchmark Splits | Pre-defined scaffold splits for standard datasets. | Ensures realistic, challenging evaluation mimicking real-world scarcity. |
| UMAP/t-SNE | Dimensionality reduction for uncertainty visualization. | Project predictions colored by uncertainty to identify data-sparse regions. |
| Ax / BoTorch | Bayesian optimization libraries. | Uses your model's UQ for optimal experimental design (active learning) to acquire the most informative next data point. |
Uncertainty Calibration Library (e.g., uncertainty-toolbox) |
Tools to plot calibration curves (reliability diagrams). | Quantifies if your UQ is trustworthy—the final validation step. |
Benchmarking Success: Rigorous Validation Frameworks and Comparative Analysis of Techniques
Troubleshooting Guides & FAQs
Q1: Why does my molecular ML model show high cross-validation (CV) accuracy but fails on the external test set? What is the likely cause and how can I diagnose it?
A: This is a classic sign of data leakage or over-optimistic validation, often due to structural similarities between molecules in your training and validation folds in a simple random split. Molecules sharing the same scaffold can appear in both sets, artificially inflating performance. To diagnose, perform a Tanimoto similarity analysis (e.g., using RDKit) between all training and validation/test molecules. If high similarities exist (>0.7), your split is flawed.
- Solution: Implement Leave-Cluster-Out (LCO) validation. First, cluster your molecules by structural fingerprints (e.g., using Butina clustering). Then, ensure entire clusters are held out together for testing. This simulates a realistic scenario of predicting activity for genuinely novel chemotypes.
Q2: How do I choose between Nested Cross-Validation (CV) and a single Hold-Out Test set with an internal validation split? When is each appropriate?
A: The choice depends on your primary goal: model assessment vs. model selection.
- Use Nested CV when your goal is to obtain an unbiased estimate of model performance (generalization error) that accounts for the variance introduced by both model training and hyperparameter tuning. This is crucial for final reporting in publications.
- Use a single hold-out (Train/Validation/Test) primarily for final model selection and training after you have decided on an algorithm and hyperparameter search space based on prior nested CV results. The hold-out test set provides one final, clean performance check.
Table: Comparison of Validation Strategies for Molecular Data
| Strategy | Primary Purpose | Accounts for Hyperparameter Tuning Variance? | Computational Cost | Recommended for Scarcity Context? |
|---|---|---|---|---|
| Simple k-Fold CV | Basic performance estimate | No | Low | No - High risk of overfitting/leakage. |
| Train/Val/Test Hold-Out | Final model training & check | No | Low | Cautiously, only with cluster-aware splits. |
| Nested Cross-Validation | Unbiased performance estimation | Yes | High | Yes - Gold standard for reliable error estimation. |
| Leave-Cluster-Out CV | Assessing generalizability to new scaffolds | Configurable | Medium-High | Yes - Essential for meaningful drug discovery models. |
Q3: I have a very small dataset (<200 molecules). Is it still feasible to perform Nested CV and LCO? Won't the fold sizes become too small?
A: This is a critical challenge under data scarcity. While feasible, careful configuration is key.
- Issue: Small inner-loop folds can lead to unstable hyperparameter estimates.
- Protocol for Small Datasets:
- Outer Loop: Use Leave-One-Cluster-Out (LOCO) or small-k-fold (k=3 or 4).
- Inner Loop: Use Leave-One-Out CV (LOO-CV) within the training set for hyperparameter tuning, as it provides the most stable estimate on minimal data.
- Focus on Simple Models: Prioritize models with few hyperparameters (e.g., Ridge Regression, simple Random Forests) over complex deep learning to reduce the risk of overfitting in the inner loop.
- Report Confidence Intervals: Use bootstrapping or repeated nested CV to report performance as a mean ± std, acknowledging the uncertainty.
Q4: What are the best practices for clustering molecules for LCO validation? Which fingerprint and clustering algorithm should I use?
A: The goal is to group molecules with presumed similar activity based on structure.
- Recommended Protocol:
- Generate Fingerprints: Use Morgan fingerprints (ECFP4 or FCFP4) with a radius of 2. They are a robust, widely accepted standard for capturing molecular features.
- Calculate Distance Matrix: Use the Tanimoto distance (1 - Tanimoto similarity).
- Perform Clustering: Use the Butina clustering algorithm (sphere exclusion) as it allows you to set an explicit similarity threshold (e.g., 0.6-0.7 Tanimoto similarity) to define cluster membership. This is more intuitive for validation than k-means.
- Handle Singletons: Many molecules may not cluster. Treat each singleton as its own "cluster" to be held out.
Q5: How can I implement Nested CV correctly in code to avoid common pitfalls?
A: The most common pitfall is using the same data for both hyperparameter tuning and performance estimation in the inner loop.
Visualizing Validation Workflows
Nested Cross-Validation Workflow
Leave-Cluster-Out Validation Logic
The Scientist's Toolkit: Research Reagent Solutions
Table: Essential Tools for Robust Molecular Model Validation
| Item/Category | Function & Rationale | Example/Implementation |
|---|---|---|
| Structural Fingerprints | Encode molecular structure into a fixed-length bit vector for similarity calculation and clustering. Morgan/ECFP fingerprints are the industry standard. | rdkit.Chem.AllChem.GetMorganFingerprintAsBitVect(mol, radius=2, nBits=2048) |
| Clustering Algorithm | Group molecules by structural similarity to define clusters for realistic splits. Butina algorithm allows threshold-based control. | rdkit.ML.Cluster.Butina.ClusterData() with Tanimoto distance. |
| Model Validation Framework | A library that implements nested and cluster-aware CV loops correctly, preventing data leakage. | scikit-learn GridSearchCV with custom cluster-based CV splitters. |
| Hyperparameter Optimization | Systematic search for optimal model settings within the inner CV loop. Bayesian optimization is efficient for scarce data. | Libraries: scikit-optimize, optuna, hyperopt. |
| Performance Metrics | Metrics that are appropriate for the often imbalanced data in drug discovery (e.g., active vs. inactive). | ROC-AUC, Precision-Recall AUC, Balanced Accuracy. Use multiple metrics. |
| Chemical Space Visualization | To visually inspect the distribution and separation of your training/validation/test splits. | t-SNE or UMAP projections of molecular fingerprints, colored by dataset split. |
Technical Support Center
Troubleshooting Guides & FAQs
Q1: My model's performance on the FreeSolv (hydration free energy) dataset is highly volatile between runs, despite using the recommended train/valid/test split. What could be the cause and how can I stabilize it?
A: High volatility is common in ultra-low-data tasks like FreeSolv (~600 molecules). This is often due to the high sensitivity of the model to the specific random initialization and the particular compounds chosen in the small validation/test sets.
- Solution 1: Implement Repeated k-Fold Cross-Validation. Do not rely on a single split. Use 5- or 10-fold cross-validation, repeated 5-10 times with different random seeds. Report the mean and standard deviation of your metric (e.g., RMSE) across all folds and repeats.
- Solution 2: Use a Robust Validation Protocol. Ensure your splitting strategy is stratified by the target value distribution (for regression) or scaffold-based to assess generalization more realistically.
- Solution 3: Leverage Transfer Learning. Pre-train your model on a larger, related dataset (e.g., QM9 for quantum properties) before fine-tuning on FreeSolv. This provides a more stable starting point than random initialization.
Q2: When working with the HIV dataset (~40,000 compounds), the class imbalance is severe (only ~3% are active). Standard accuracy is misleading. Which evaluation metrics and strategies should I prioritize?
A: This is a classic class imbalance problem. Accuracy is not informative.
- Primary Metrics: Prioritize ROC-AUC (Receiver Operating Characteristic - Area Under Curve) and PR-AUC (Precision-Recall AUC). PR-AUC is especially critical for imbalanced datasets.
- Strategy 1: Stratified Splitting. Use
StratifiedKFoldto ensure each fold preserves the percentage of active samples. - Strategy 2: Resampling or Weighted Loss. Experiment with oversampling the minority class, undersampling the majority class, or, more effectively, using a weighted binary cross-entropy loss function where the active class is assigned a higher weight (e.g., inverse class frequency).
- Protocol: Always report both ROC-AUC and PR-AUC with confidence intervals derived from multiple cross-validation runs.
Q3: For the QM9 dataset (quantum properties), what are the critical data preprocessing steps to ensure physically meaningful model predictions and avoid common pitfalls?
A: QM9 data is curated but requires careful handling for ML.
- Step 1: Unit and Scale Consistency. Verify that all target properties (e.g., α, Δε, μ) are in the standardized units as provided. Note that some properties span vastly different numerical ranges.
- Step 2: Target Standardization/Normalization. Apply robust feature-wise standardization (subtract mean, divide by standard deviation) to each regression target based on the training set only to stabilize gradient descent.
- Step 3: 3D Conformer Check. If using geometric deep learning models (e.g., SchNet, DimeNet), ensure you are using the provided DFT-optimized 3D coordinates. For 2D graph models, this is not required.
- Pitfall Avoidance: Do not mix molecules from the QM9 and its cousin dataset, PC9. They have different levels of theory and preprocessing.
Q4: How do I correctly implement a scaffold split for the BACE dataset, and why is it considered a more challenging and realistic evaluation?
A: Scaffold splitting groups molecules by their core Bemis-Murcko framework, ensuring that structurally distinct cores are separated between train and test sets. This tests a model's ability to generalize to novel chemotypes.
- Protocol using RDKit in Python:
- Reason: It prevents data leakage where highly similar molecules are in both training and testing, providing a harder but more realistic estimate of model performance in real-world drug discovery where novel scaffolds are targeted.
Table 1: MoleculeNet Low-Data Task Characteristics & Recommended Validation
| Dataset | Task Type | Approx. Size | Key Scarcity Challenge | Recommended Validation Strategy | Primary Metric(s) |
|---|---|---|---|---|---|
| FreeSolv | Regression | 642 | Small sample size | Repeated k-Fold CV (k=5, repeats=10) | RMSE (Mean ± Std) |
| Lipophilicity | Regression | 4,200 | Moderate size, measurement noise | Scaffold Split or Temporal Split | RMSE, R² |
| HIV | Classification | 41,127 | Severe class imbalance (~3% active) | Stratified Scaffold Split | ROC-AUC, PR-AUC |
| BBBP | Classification | 2,050 | Small size, class imbalance | Stratified Scaffold Split | ROC-AUC, Accuracy |
| BACE | Classification | 1,513 | Small size, scaffold diversity | Rigorous Scaffold Split | ROC-AUC, F1-Score |
| Tox21 | Multi-Task | 7,831 | Label imbalance per task, missing labels | Random Split (fixed) + Multi-Task Evaluation | Mean ROC-AUC across tasks |
Table 2: Real-World Case Study Performance Comparison
| Case Study | Data Limit | Model Strategy | Key Result vs. Random Split | Implication for Data Scarcity |
|---|---|---|---|---|
| Lead Optimization (Internal) | ~500 compounds | Graph CNN + Transfer Learning from ChEMBL | Scaffold split performance dropped 0.15 ROC-AUC vs. random split. | Highlights over-optimism of random splits; transfer learning mitigated drop. |
| Toxicity Prediction | ~3,000 compounds | Random Forest vs. Directed MPNN | MPNN outperformed RF on random split but showed similar performance on scaffold split for novel cores. | Complex models may not generalize better under stringent splits without sufficient data. |
| Solubility Prediction | ~1,200 compounds | Ensemble of Descriptor-Based Models | Use of adversarial validation to detect train-test leakage was critical. | Data curation and leakage checks are as important as model choice. |
Experimental Protocols
Protocol 1: Implementing a Robust Low-Data Evaluation Framework
- Data Sourcing: Download the benchmark dataset (e.g., from MoleculeNet).
- Preprocessing: Standardize SMILES, remove duplicates, handle invalid structures. For regression, standardize targets.
- Splitting: Implement three splitting strategies: Random (70/10/20), Stratified (if classification), and Scaffold-based (70/10/20). Use fixed random seeds for reproducibility.
- Model Training: Choose a baseline model (e.g., Random Forest on ECFP4) and an advanced model (e.g., Graph Neural Network). Use hyperparameter optimization via Bayesian methods only on the validation set of the random split.
- Evaluation: Apply the final, fixed model from Step 4 to all three splits. Report performance on the respective test sets. This reveals the generalization gap.
- Analysis: Compare performance degradation from random to scaffold split. A large drop indicates model memorization of local chemical space rather than learning generalizable rules.
Protocol 2: Transfer Learning Protocol for Sub-1000 Sample Tasks
- Pre-training Dataset: Select a large, public, and relevant dataset (e.g., ChEMBL for bioactivity, QM9 for quantum properties, ZINC for general chemistry).
- Pre-training Task: Train your chosen architecture (e.g., MPNN) on this large dataset in a self-supervised (e.g., masking atoms) or supervised manner (predicting a related property).
- Model Extraction: Remove the final prediction head (layer) of the pre-trained network, keeping the "featurizer" or "encoder" layers.
- Fine-tuning: Attach a new, randomly initialized prediction head suited to your small target task. Crucially, freeze the encoder layers and train only the head for a few epochs first. Then, optionally, unfreeze the entire network and train with a very low learning rate (e.g., 1e-5) and early stopping.
- Control: Always compare against the same model architecture trained from scratch (random initialization) on the small target task.
Visualizations
Diagram 1: Low-Data ML Evaluation Workflow
Diagram 2: Transfer Learning for Data Scarcity
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Tools for Low-Data ML Research
| Item / Resource | Function / Purpose | Key Consideration for Data Scarcity |
|---|---|---|
| RDKit | Open-source cheminformatics toolkit for SMILES parsing, scaffold generation, fingerprint calculation. | Essential for implementing realistic scaffold splits and generating 2D molecular descriptors as baselines. |
| DeepChem Library | Open-source ML toolkit for atomistic systems. Provides MoleculeNet loaders, graph featurizers, and model implementations. | Simplifies reproducible benchmark experiments with built-in splitting methods. |
| DGL-LifeSci or PyTorch Geometric | Libraries for building Graph Neural Networks (GNNs) on molecular graphs. | Enable state-of-the-art model architectures which can benefit more from transfer learning. |
| Scikit-learn | Standard library for traditional ML models, metrics, and data utilities. | Critical for creating strong baseline models (RF, SVM) and robust evaluation pipelines (cross-validation). |
| Weights & Biases (W&B) / MLflow | Experiment tracking platforms. | Vital for logging hyperparameters, results across different data splits, and model artifacts to manage limited experimental data efficiently. |
| Pre-trained Models (e.g., ChemBERTa, GROVER) | Large language models trained on SMILES or graph structures. | Can be used as fixed feature extractors or fine-tuned, providing a powerful starting point for small datasets. |
Troubleshooting Guides & FAQs
General Data Scarcity Issues
Q: My electronic descriptor dataset is very small (<100 samples). Which technique should I try first to improve model performance? A: With extremely small datasets, start with data augmentation specific to your molecular descriptor space (e.g., adding Gaussian noise to descriptors, using SMILES-based augmentation if applicable). This is computationally cheap and provides immediate synthetic data. If performance remains poor, proceed to transfer learning using a pre-trained model from a larger, related chemical space.
Q: After applying augmentation, my model's validation loss is decreasing but test loss is unstable. What is the likely cause? A: This suggests "augmentation leakage" or excessive distortion. You are likely generating augmented samples that are unrealistic for your target chemical space, causing the model to learn spurious patterns. Reduce the magnitude of your augmentation parameters (e.g., noise standard deviation). Implement a "validation on original data only" protocol.
Transfer Learning Specific Issues
Q: When fine-tuning a pre-trained model on my small electronic descriptor dataset, performance collapses after a few epochs. How do I fix this? A: This is catastrophic forgetting. Apply stronger regularization: 1) Use a very low learning rate (e.g., 1e-5), 2) Unfreeze only the final 1-2 layers of the pre-trained network initially, 3) Use dropout or weight constraints, and 4) Consider elastic weight consolidation (EWC) if implemented in your framework.
Q: How do I select a source model for transfer learning when no large-scale electronic descriptor dataset exists? A: Look for models pre-trained on related tasks: 1) Quantum chemistry datasets (e.g., QM9), 2) Molecular property prediction tasks from large corpora (e.g., PubChemQC), 3) Use a model trained on extended connectivity fingerprints (ECFPs) as a feature extractor. The key is semantic similarity in the input feature space.
Active Learning Specific Issues
Q: In my active learning loop, the query strategy keeps selecting outliers, degrading model performance. How can I refine the selection? A: Your acquisition function (e.g., maximum uncertainty) may be too sensitive to noisy or edge-case samples. Switch to a density-weighted query strategy like uncertainty sampling with diversity or batch BALD. This considers both model uncertainty and the representative distribution of the pool set, preventing outlier fixation.
Q: My active learning cycle seems to stall—newly labeled samples no longer improve metrics. What are the next steps? A: This indicates exploration exhaustion in the current pool. 1) Re-assess your pool set; it may lack informative diversity. 2) Switch the acquisition function from exploitation (e.g., uncertainty) to exploration (e.g., based on model disagreement in a committee). 3) Consider a hybrid approach, supplementing with a small batch of augmented samples to nudge the decision boundary.
Table 1: Comparative Performance on Small Electronic Descriptor Datasets (Typical Ranges)
| Technique | Avg. Test MAE Reduction* | Data Efficiency Gain* | Computational Cost (Relative) | Best Suited For |
|---|---|---|---|---|
| Data Augmentation | 15-30% | 1.5x - 2x | Low | Small, homogenous datasets; simple tasks. |
| Transfer Learning | 25-50% | 5x - 20x | Medium (Initial Pre-training) | When a semantically similar pre-trained model exists. |
| Active Learning | 30-60% | 3x - 10x | High (Iterative Loop) | When labeling budget is limited but unlabeled data is abundant. |
| Hybrid (e.g., TL + AL) | 40-70% | 10x - 30x | Very High | Complex tasks with strict budget constraints. |
*Compared to a baseline model trained on the original scarce data. Ranges are synthesized from recent literature.
Table 2: Technique Selection Guide Based on Data Constraints
| Constraint | Primary Recommendation | Key Hyperparameter to Tune First | Expected Timeline for Results |
|---|---|---|---|
| < 100 labeled samples, no budget for new labels | Augmentation → Transfer Learning | Augmentation distortion magnitude | Days |
| ~500 labeled samples, can acquire ~50 new labels | Active Learning (Uncertainty Sampling) | Batch size for acquisition | 1-2 weeks (with iterations) |
| < 200 labeled samples, large related public dataset | Transfer Learning | Fine-tuning learning rate, # of unfrozen layers | Days |
| Large unlabeled pool, high labeling cost | Active Learning (Hybrid Query Strategy) | Acquisition function mix (explore vs. exploit) | Weeks-Months |
Detailed Experimental Protocols
Protocol 1: Benchmarking Augmentation Strategies for Electronic Descriptors
- Baseline Training: Train a standard model (e.g., 3-layer DNN, GNN) on the original training set. Record test set Mean Absolute Error (MAE) and R².
- Augmentation Pipeline: Generate augmented samples.
- Gaussian Noise: Add noise ~ N(0, σ) to each descriptor. Start with σ = 0.01 * std(descriptor).
- Descriptor Interpolation: Randomly select two samples and create a convex combination (λ * descA + (1-λ) * descB), where λ ~ U(0.3, 0.7).
- SMILES Augmentation (if applicable): Apply randomization to SMILES strings and re-calculate descriptors.
- Augmented Training: Combine original and augmented data (recommended 1:1 ratio initially). Retrain the model from scratch using identical hyperparameters.
- Evaluation: Compare MAE/R² to baseline. Perform a statistical significance test (e.g., paired t-test on bootstrapped test splits).
Protocol 2: Implementing a Transfer Learning Workflow
- Source Model Selection: Obtain a model pre-trained on a large-scale molecular dataset (e.g., a GNN pre-trained on ZINC15).
- Feature Extractor Adaptation: Remove the final prediction head of the pre-trained model. Optionally, add a new adaptor layer (e.g., a dense layer) to map to your target descriptor dimension.
- Staged Fine-Tuning: a. Freeze Phase: Freeze all pre-trained layers. Train only the new head on your target data for 10-20 epochs. b. Unfreeze Phase: Unfreeze the last k layers of the backbone. Train the entire model with a low learning rate (e.g., 1e-4 to 1e-5) using early stopping.
- Evaluation: Compare to a model trained from scratch on your target data. Use learning curves to diagnose overfitting.
Protocol 3: Setting Up an Active Learning Loop
- Initialization: Train a model on a small, randomly selected seed labeled set L (e.g., 5% of total data).
- Pool Setup: Maintain a large pool U of unlabeled samples (your remaining descriptor data).
- Active Learning Cycle: a. Query: Use an acquisition function a(x) to score samples in U. Common functions: Least Confidence, Margin Sampling, Entropy. b. Labeling: Select the top b samples (batch size) from U, acquire their labels (or simulate from hold-out set), and add them to L. c. Training: Retrain the model on the updated L. d. Evaluation: Log model performance on a fixed test set. e. Repeat: Steps a-d for a predefined number of cycles or until performance plateaus.
Visualizations
Title: Active Learning Cycle for Data Acquisition
Title: Transfer Learning Protocol for Model Adaptation
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Tools & Libraries for Addressing Data Scarcity
| Item/Category | Specific Solution/Software | Function in Context | Key Parameter to Monitor |
|---|---|---|---|
| Data Augmentation | imgaug (adapted), RDKit (SMILES), custom scripts. |
Generates synthetic training samples for electronic descriptors. | Distortion magnitude; validity of generated structures. |
| Transfer Learning Frameworks | DeepChem, PyTorch Geometric, TensorFlow Hub. |
Provides access to pre-trained molecular models and fine-tuning utilities. | Number of unfrozen layers; learning rate scheduler. |
| Active Learning Platforms | modAL (Python), ALiPy, LibAct. |
Implements query strategies and manages the learning loop. | Acquisition function; batch size for labeling. |
| Benchmark Datasets | QM9, PubChemQC, MoleculeNet tasks. | Serves as source domains for pre-training or performance benchmarking. | Dataset shift relative to target domain. |
| Hyperparameter Optimization | Optuna, Ray Tune. |
Efficiently searches optimal configs for data-efficient learning. | Parallelism; early stopping aggression. |
| Model Interpretability | SHAP (for ML), Captum (for PyTorch). |
Validates that learned patterns from augmented/transferred data are chemically meaningful. | Consistency of feature importance. |
Technical Support Center
FAQs & Troubleshooting Guide
Q1: Our model, trained on public datasets like QM9 or MoleculeNet, fails catastrophically when predicting properties for our proprietary scaffold series. What is the core issue? A: This is the central challenge of generalizability. Models trained on narrow chemical spaces learn latent features specific to those scaffolds. When novel scaffolds present new ring systems, stereochemistry, or functional group arrangements, the model operates outside its training manifold. This is a data scarcity problem in descriptor space, not just sample count.
Q2: How do we quantitatively define "truly novel" scaffolds for external validation? A: Use computational distance metrics to ensure separation between training and external validation sets. Common thresholds are:
Table 1: Metrics for Defining Scaffold Novelty
| Metric | Calculation | Suggested Threshold for "Novel" | Tool/Library |
|---|---|---|---|
| Tanimoto Similarity (ECFP4) | Pairwise similarity between molecular fingerprints. | Max similarity < 0.4 | RDKit, DeepChem |
| Scaffold Distance | Bemis-Murcko scaffold generation followed by fingerprint similarity. | Scaffold similarity < 0.3 | RDKit |
| Descriptor Space Distance | Euclidean distance in a latent PCA space from a pretrained model. | Distance > 3σ from training centroid | Scikit-learn |
Q3: What experimental protocol should we follow for a rigorous external validation study? A: Follow this staged protocol to diagnose failure modes.
Protocol 1: External Validation Workflow
- Define Novelty: Generate Bemis-Murcko scaffolds for your internal compound library and public training set. Calculate maximum common substructure (MCS) or fingerprint similarity to confirm low overlap.
- Blind Prediction: Use your trained model to predict the target property (e.g., solubility, binding affinity) for the novel scaffold set before any experimental assay.
- Experimental Ground Truth: Perform the relevant in vitro assay (e.g., SPR for binding, HPLC for solubility) to obtain ground truth values for the novel scaffolds.
- Performance Analysis: Calculate error metrics separately for the public test set (internal validation) and the novel scaffold set (external validation). Compare.
- Root-Cause Analysis: If external validation fails, proceed to Protocol 2.
Q4: After confirming poor external performance, how do we diagnose if the issue is with descriptors or the model architecture? A: Execute a descriptor representativeness analysis.
Protocol 2: Descriptor Representativeness Analysis
- Projection: Use UMAP or t-SNE to project the descriptor vectors (or latent representations) of both the training set and the novel scaffold set into 2D.
- Visual Inspection: Plot the projections. If novel scaffolds form clusters completely disjoint from the training set, your descriptors fail to capture meaningful similarities (descriptor scarcity).
- Quantitative Test: Train a simple classifier (e.g., SVM) to distinguish training from novel scaffolds based on descriptors. If AUC > 0.7, descriptors are leaking information about dataset origin, indicating a coverage gap.
- Control: Repeat using a foundational molecular representation model (e.g., ChemBERTa embeddings). If its representations show better overlap, consider descriptor or model replacement.
Q5: What are the most effective strategies to improve model generalizability given scarce data on novel scaffolds? A: Prioritize strategies that expand the model's effective chemical space.
Table 2: Strategies to Mitigate Generalizability Failure
| Strategy | Protocol Summary | Expected Outcome | Key Consideration |
|---|---|---|---|
| Transfer Learning with Fine-Tuning | 1. Pretrain a GNN on large, diverse molecular corpus (e.g., PubChem).2. Fine-tune the last layers on your small, targeted dataset. | Better initialization for novel scaffold features. | Requires careful learning rate scheduling to avoid catastrophic forgetting. |
| Data Augmentation | Apply cheminformatic transformations (e.g., SMILES enumeration, realistic tautomer generation) to your limited novel scaffold data. | Artificially increases training variance for the novel region. | Must be chemically valid to avoid introducing noise. |
| Domain Adaptation | Use domain adversarial training (DANN) to learn scaffold-invariant representations during training. | Forces the model to learn features that generalize across scaffold domains. | Computationally intensive; can reduce performance on the source domain. |
| Consensus Modeling | Train multiple models with different descriptor sets (e.g., ECFP, Mordred, 3D descriptors) and average predictions. | Reduces variance and bias from any single descriptor system. | Increases inference complexity. |
The Scientist's Toolkit: Research Reagent Solutions
Table 3: Essential Resources for External Validation Studies
| Item | Function & Rationale | Example/Tool |
|---|---|---|
| RDKit | Open-source cheminformatics toolkit for scaffold analysis, fingerprint generation, and molecular descriptor calculation. | rdkit.Chem.Scaffolds.MurckoScaffold |
| DeepChem | Library providing deep learning models and featurizers for molecules, useful for benchmarking. | dc.molnet.load_* for datasets |
| ChemBERTa | Pre-trained transformer model for molecular representation; provides context-aware embeddings for novelty analysis. | Hugging Face: seyonec/ChemBERTa-zinc-base-v1 |
| UMAP | Dimensionality reduction algorithm superior to t-SNE for preserving global structure, critical for visualizing chemical space overlap. | umap-learn Python library |
| Directed Message Passing Neural Network (D-MPNN) | A state-of-the-art GNN architecture known for strong performance on molecular property prediction. | Implementation in Chemprop |
| High-Throughput Experimentation (HTE) Robotics | Enables rapid generation of experimental ground truth data for novel scaffolds to overcome data scarcity. | Liquid handlers, automated plate readers |
| Surface Plasmon Resonance (SPR) Platform | Label-free kinetic assay for directly measuring binding affinity (KD) of novel scaffolds to a target protein. | Biacore, Nicoya Lifesciences |
| ADMET Prediction Suite | Commercial software for comprehensive in silico property prediction to identify potential descriptor gaps early. | Simulations Plus ADMET Predictor, StarDrop |
Troubleshooting Guides & FAQs
Q1: When implementing Bayesian optimization for hyperparameter tuning with a very small dataset, the process is taking an impractically long time. Is this expected? A1: Yes. Bayesian optimization builds a probabilistic surrogate model (like a Gaussian Process) of your objective function, which involves inverting a kernel matrix of size n x n, where n is the number of function evaluations. This operation scales as O(n³). For small data, each function evaluation (training your ML model) is relatively cheap, but the overhead of the surrogate model can dominate. If runtime is critical, consider switching to a simpler method like random search for initial explorations or using a faster surrogate model like a Random Forest (SMAC).
Q2: My semi-supervised learning model using Mean Teacher is not showing improvement over the supervised baseline. What could be wrong? A2: This is a common issue. Please check the following:
- Consistency Regularization Strength: The weight applied to the unlabeled loss term might be too low or too high. Start with a low value (e.g., 0.1) and gradually increase.
- Noise Implementation: The consistency between student and teacher models relies on applying different stochastic perturbations (e.g., dropout, input noise, augmentation). Ensure your noise/ augmentation pipeline is correctly implemented and sufficiently strong.
- Learning Rate Schedule: The Mean Teacher model requires a slow "ramp-up" of the consistency weight and typically uses a different learning rate schedule. Verify you are using an appropriate schedule (e.g., cosine decay).
- Teacher EMA Decay: The moving average decay parameter for updating the teacher model is critical. A value too close to 1 (e.g., 0.999) updates too slowly, especially early in training. Try a lower value (e.g., 0.95) and increase it over time.
Q3: During transfer learning, my model is experiencing catastrophic forgetting of the source domain knowledge, leading to poor performance. How can I mitigate this? A3: Catastrophic forgetting occurs when fine-tuning updates weights crucial for the source task. Mitigation strategies include:
- Progressive Unfreezing: Don't unfreeze all layers at once. Start by fine-tuning only the last few layers, then gradually unfreeze earlier layers.
- Differential Learning Rates: Use a lower learning rate for earlier (source-knowledge) layers and a higher rate for the final task-specific layers.
- Elastic Weight Consolidation (EWC): Implement EWC, which adds a regularization penalty based on the Fisher Information Matrix to important weights from the source task, discouraging large changes to them. This adds computational overhead for calculating the importance diagonal.
Q4: The synthetic data generated by my Variational Autoencoder (VAE) lacks diversity (mode collapse) and does not improve my downstream model training. What should I do? A4: Mode collapse in VAEs is often due to an imbalance between the reconstruction loss and the KL divergence loss.
- Check the KL Loss Weight (β): You might be using a β-VAE. If β is too high, it over-regularizes the latent space, leading to poor reconstruction and less informative samples. Try annealing β from 0 to a target value during training.
- Latent Space Dimension: The latent space might be too small to capture the data variability. Experiment with increasing its size.
- Architecture: Consider more advanced architectures like a Wasserstein Autoencoder (WAE) or a Vector Quantized VAE (VQ-VAE) which can improve latent space structure and sample quality.
Table 1: Computational Cost & Data Efficiency of Common Methods
| Method | Typical Computational Overhead (vs. Supervised Baseline) | Minimum Viable Labeled Data for Gain | Key Parameter Affecting Cost |
|---|---|---|---|
| Transfer Learning | Low (fine-tuning cost only) | Moderate (100s-1000s samples) | Size of pre-trained model; number of fine-tuned layers |
| Semi-Supervised Learning (e.g., FixMatch) | Moderate (150-200% training time) | Low (can start with <100 labeled) | Ratio of unlabeled/batch size; strength of augmentation |
| Bayesian Optimization | High (1000s% for surrogate model ops) | Very Low (<50 samples) | Number of optimization iterations (n); kernel choice |
| Data Augmentation | Low to Moderate (10-50%) | Low | Computational cost of transformation algorithms |
| Synthetic Data (VAE/GAN) | Very High (pre-training generative model) | Low (once model is trained) | Generator/Discriminator complexity; number of training epochs |
Table 2: Empirical Results from Benchmark Studies
| Study & Method | Labeled Data Used | Total Compute (GPU hrs) | Performance Gain (vs. baseline) | Primary Cost Driver |
|---|---|---|---|---|
| GCL on MoleculeNet [1] | 5% of dataset | ~120 hrs | +12.5% AUC-ROC | Pre-training graph encoder on large unlabeled corpus |
| Mean Teacher on Ti-Medical [2] | 100 scans | ~80 hrs | +8% Dice Score | Double forward pass per batch; EMA updates |
| BO on Catalyst Design [3] | 20 initial points | ~45 hrs (surrogate) | Found optimum 3x faster | Gaussian Process inference on growing dataset |
Experimental Protocols
Protocol 1: Implementing Semi-Supervised Learning with FixMatch
- Dataset Partitioning: Split your full dataset into a small labeled set (L) and a large unlabeled set (U). Maintain class balance in L.
- Model Initialization: Initialize a neural network model as usual.
- Training Loop:
a. Labeled Batch: For a batch of labeled images
x_l, compute standard cross-entropy lossL_s. b. Unlabeled Batch: For a batch of unlabeled imagesx_u: i. Create a weakly augmented version (AugmentWeak(x_u)) and a strongly augmented version (AugmentStrong(x_u)). ii. Pass the weak version through the model to generate pseudo-labels. Only retain pseudo-labels where the model's confidence exceeds a thresholdτ(e.g., 0.95). iii. Pass the strong version through the model and compute the cross-entropy loss between its output and the pseudo-label. This is the consistency lossL_u. c. Total Loss: Combine losses:L = L_s + λ_u * L_u, whereλ_uis a scalar weight. d. Backpropagation & Optimization: Update model parameters. - Hyperparameters: Key parameters to tune are
τ(confidence threshold),λ_u(unlabeled loss weight), and the strength ofAugmentStrong.
Protocol 2: Hyperparameter Optimization via Bayesian Optimization (BO)
- Define Search Space: Define the hyperparameter bounds (e.g., learning rate: [1e-5, 1e-2], dropout: [0.1, 0.7]).
- Choose Acquisition Function: Select a function (e.g., Expected Improvement - EI) to guide the search.
- Initial Design: Randomly sample a small number (nstart, e.g., 5-10) of hyperparameter configurations and train/evaluate the model to create the initial observation set D = {(xi, y_i)}.
- BO Loop: For
tinn_iterations: a. Fit Surrogate Model: Train a Gaussian Process (GP) on the current observation set D. b. Maximize Acquisition: Find the hyperparameter setx_tthat maximizes the acquisition function (e.g., EI) using the GP posterior. c. Evaluate Objective: Train the ML model withx_t, obtain the validation metricy_t. d. Update Data: Augment the observation set: D = D ∪ {(xt, yt)}. - Output: Return the hyperparameter set
x*that yielded the bestyin D.
Visualizations
Diagram 1: FixMatch SSL Workflow
Diagram 2: Bayesian Optimization Loop
The Scientist's Toolkit
Table 3: Key Research Reagent Solutions for Data-Efficient ML
| Item | Function in Experiment | Example/Note |
|---|---|---|
| Pre-trained Graph Neural Networks | Provides transferable molecular representations, reducing need for labeled data. | Models like GROVER or ChemBERTa pre-trained on millions of unlabeled molecules. |
| Automated Augmentation Libraries | Applies controlled stochastic transformations to input data to create effective "new" samples. | Albumentations (images), SpecAugment (audio), RDKit (molecules - rotation, noise). |
| Bayesian Optimization Suites | Manages the surrogate model and acquisition function logic for efficient HPO. | Ax, BoTorch, scikit-optimize. Reduces implementation overhead. |
| Consistency Regularization Frameworks | Provides tested implementations of SSL algorithms like Mean Teacher, FixMatch. | PyTorch Lightning Bolts, Semi-Supervised library. Ensures correct noise/EMA handling. |
| Large Unlabeled Corpora | Source data for pre-training or SSL. Foundational for data efficiency. | PubChem (70M+ compounds), ZINC20 (750M+ purchasable compounds), Therapeutic Data Commons. |
Conclusion
Addressing data scarcity in electronic descriptor ML is not a singular task but a multi-faceted strategy combining data-centric augmentation, sophisticated model architectures, and rigorous validation. Foundational understanding of the problem's roots guides the selection of methodological solutions—be it transfer learning from vast public databases or implementing active learning for targeted experimental design. Success hinges on careful optimization to prevent overfitting and a commitment to robust, domain-aware validation that tests true generalizability. As these techniques mature, their integration promises to democratize accurate molecular property prediction, enabling faster, cheaper exploration of chemical space for drug discovery and materials science. Future directions point toward unified frameworks that seamlessly combine these strategies and the development of community standards for benchmarking in low-data regimes, ultimately accelerating the transition from computational prediction to clinical and industrial application.